Original research |
Peer reviewed |
Statistical process control methods used to evaluate the serologic responses of pigs infected with three Salmonella serovars
Métodos de control estadístico del proceso utilizado para evaluar las respuestas serológicas de cerdos infectados con tres serovariedades de Salmonella
Méthodes statistiques de contrôle du processus utilisées pour évaluer les réponses sérologiques de cochons infectés avec trois serovars de Salmonella
D. H. Baum, MS, DVM, PhD; S. Ward, PhD; C. L. Baum, MS; N. Lee; D. D. Polson, DVM, MS, PhD; D. L. Harris, DVM, PhD; B. Nielsen, DVM, PhD
DHB, DDP: Boehringer Ingelheim Vetmedica Inc, Ames, Iowa. SW: Statistical Process Controls Inc, Knoxville, Tennessee. CLB, NL, DLH: Department of Microbiology, Iowa State University, Ames, Iowa. DLH: Department of Veterinary Diagnostics and Production Animal Medicine, Iowa State University, Ames, Iowa. BN: Danish Bacon and Meat Council, Axelborg, Copenhagen V, Denmark. Corresponding author: Dr David H. Baum, Circle Four Farms, PO Box 100, Milford, UT 84751; E-mail: dbaum@c4farms.com.
Cite as: Baum DH, Ward S, Baum CL, et al. Statistical process control methods used to evaluate the serologic responses of pigs infected with three Salmonella serovars. J Swine Health Prod. 2005;13(6):304-313.
Also available as a PDF.
SummaryObjectives: To confirm that the mix-ELISA detects antibody against Salmonella serovars Typhimurium, Infantis, and Choleraesuis; to demonstrate that statistical process control (SPC) methods can be used to validate the mix-ELISA; and to demonstrate how SPC can be used to assess the Salmonella serologic status of swine. Methods: Three groups of pigs were inoculated with Salmonella Typhimurium, Salmonella Choleraesuis, or Salmonella Infantis (one serovar per group). Serologic responses were measured with the mix-ELISA and compared to responses of a group of uninfected pigs. Mix-ELISA results were evaluated using SPC methods to calculate a positive-negative cutoff value and to determine assay diagnostic sensitivity and specificity. The SPC results were compared to results of receiver operator characteristic (ROC) curve analysis. Results: Three cutoffs were determined from the SPC methods: group average (optical density [OD]% > 7.239), group range (OD% range >= 12.61), and individual (OD% >= 12). ROC curve analysis also showed optimized sensitivity (0.845) and specificity (1.00) when the individual cutoff was OD% >= 12.5. The OD% values were highest in pigs infected with Salmonella Typhimurium. Implications: The mix-ELISA detects antibody in pigs infected with Salmonella serovars, including Choleraesuis. Statistical process control methods can be used with mix-ELISA results to determine diagnostic cutoff values for assessing Salmonella serologic status. The degree of Salmonella exposure in swine can be assessed using SPC methods. | ResumenObjetivos: Confirmar que la mix-ELISA detecta anticuerpos contra las serovariedades de Salmonella Typhimurium, Infantis, y Choleraesuis; para demostrar que el método estadístico de control del proceso (SPC por sus siglas en inglés) puede utilizarse para validar la mix-ELISA; y para demostrar cómo se puede utilizar el SPC para evaluar el estado serológico de Salmonella de los cerdos. Métodos: Se inocularon tres grupos de cerdos con Salmonella Typhimurium, Salmonella Choleraesuis ó Salmonella Infantis (una serovariedad por grupo). Se midieron las respuestas serológicas con la mix-ELISA y se compararon con las respuestas de un grupo de cerdos no infectados. Los resultados de la mix-ELISA se evaluaron utilizando métodos del SPC para calcular el punto de corte positivo-negativo y para determinar la especificidad y la sensibilidad de la prueba. Los resultados del SPC se compararon con los resultados del análisis de curva de las características del operador receptor (ROC por sus siglas en inglés). Resultados: Se determinaron tres puntos de corte a partir de los métodos SPC: promedio del grupo (densidad óptica [OD% por sus siglas en inglés] > 7.239), rango del grupo (OD% rango >= 12.61) e individual (OD% >= 12). El análisis de curva ROC también mostró una optimización en la sensibilidad (0.845) y especificación (1.00) cuando el punto de corte fue OD% >= 12.5. Los valores del OD% fueron los más altos en los cerdos infectados con Salmonella Typhimurium. Implicaciones: La mix-ELISA detecta anticuerpos en cerdos infectados con serovariedades de Salmonella, incluyendo la Choleraesuis. El método de control estadístico del proceso puede utilizarse en conjunto con los resultados de la mix-ELISA para determinar los valores de corte para valorar el estado serológico de la Salmonella. El grado de exposición de la Salmonella en los cerdos puede valorarse utilizando el método de SPC. | ResuméObjectifs: Confirmer que la mix-ELISA détecte des anticorps contre le serovars de Salmonella Typhimurium, Infantis et Choleraesuis; pour démontrer que le méthode statistique de contrôle du processus (SPC par ses sigles en anglais) peut être utilisée pour valider la mix-ELISA; et pour démontrer comme le SPC peut être utilisé pour évaluer le statut sérologique de Salmonella du porcs. Méthodes: Trois groupes de cochons ont été inoculés avec la Salmonella Typhimurium, la Salmonella Choleraesuis, ou la Salmonella Infantis (un serovar par groupe). Les réponses sérologiques ont été mesurées avec la mix-ELISA et ils ont été comparés aux réponses d'un groupe de porcs qui n'étaient pas infectés. Les résultats de la mix-ELISA ont été évalués en utilisant des méthodes SPC pour calculer la valeur limite (cutoff) positif-négative et pour déterminer la spécificité et la sensibilité diagnostic de l'épreuve. Les résultats du SPC ont été comparés aux résultats de l'analyse de courbe de la caractéristique de l'opérateur receveur (ROC par ses sigles en anglais). Résultats: Trois limites a partir des méthodes SPC ont été déterminés: moyenne du groupe (densité optique [OD% par ses sigles en anglais] > 7.239), portée du groupe (OD% portée >= 12.61), et individuel (OD% >= 12). Le analyse de courbe ROC a montré aussi la sensibilité optimisée (0.845) et la spécificité (1.00) quand le limite individuel a été OD% >= 12.5. Les valeurs du OD% ont été les plus hautes dans les porcs infectés avec Salmonella Typhimurium. Implications: La mix-ELISA détecte des anticorps dans les cochons infectés avec le serovars de Salmonella, y compris le Choleraesuis. Les méthodes statistiques de contrôle du processus peuvent être utilisées avec les résultats de la mix-ELISA pour déterminer les valeurs limites pour évaluer le statut sérologique de Salmonella. Le degré d'exposition de Salmonella dans les cochons peut être estimé en utilisant les méthodes de SPC. |
Keywords: swine, statistical
process control, Salmonella, ELISA
Search the AASV web site
for pages with similar keywords.
Received: September
12, 2002
Accepted: March
2, 2004
Different sources of variation are inherent to serologic assays. Two sources of variation are the individual animal and the group of animals being tested. The immunologic status of any individual animal or group of animals is dynamic.1 That is, the detected concentration of antibody in serum samples is likely to vary from one day to the next when the same animal is sampled and tested using the same antibody assay. There is also variation in the same sample within the same laboratory over time. That is, there is a degree of variation inherent in the assay system. Determination of cutoff values for serologic assays should account for these sources of variation.
A method that accounts for this expected variation (dispersion) of antibody concentration for known-positive and known-negative animals would allow for analytical rather than empirical assessment of diagnostic test results.2 Thus, it might be possible to determine with defined limits of certainty whether an individual animal or a group of animals is seropositive or seronegative for a given condition or pathogen, on the basis of the test result and the variation within the sampled population. Such an assessment of test results would be useful for pathogen reduction programs that address swine herd health or preharvest food safety, in the same manner as has been used for the continuous improvement of processes used in manufacturing.3-7
Optimal use of serologic testing requires a working knowledge of disease pathogenesis, immunodiagnostic techniques, and statistical inference.2 Classical validation of diagnostic tests requires testing many samples from known-infected (true-positive) and known-uninfected (true-negative) animals.8 Diagnostic sensitivity and specificity are then determined by use of a 2 x 2 table.9 The foundation for calculation of sensitivity and specificity is determination of a cutoff value for the assay.10 Cutoff values (critical values) are used to determine whether a sample is positive or negative for the condition being tested for.8 The cutoff value can be determined by several methods, for instance, empirically from histograms of true-positive and true-negative animals.11,12 Receiver operating characteristic (ROC) curve analysis uses the values from true-positive and true-negative samples to optimize diagnostic sensitivity and specificity.11 However, ROC analysis does not account for within-group variation. Assays can be compared by the respective areas under their ROC curves.11 Greiner et al11 described other methods for determining cutoff values, none of which account for the variation that may exist in the tested populations. In addition, these methods tend to determine cutoff values for individuals, not groups (ie, herd assays). The Gaussian (normal) distribution has been used to determine cutoff values, but is not an adequate method for skewed or multimodal data distributions.11
Statistical process control (SPC) methods were developed by Shewhart3,4 to account for the variation that exists within k subgroups of n observations. Over time, SPC permits each subgroup's variation differences to be assessed.3,6 The four foundations of SPC charts are as follows.13 First, control limits are always set at a distance of three sigma units on either side of the central line.13 Second, sigma has been defined as the standard deviation (SD) of a homogeneous set of data and is estimated using an average dispersion statistic (the average range) or a median dispersion statistic (the median range).13 Sigma is used in the context of the empirical rule6 applied to a homogeneous set of data: approximately 60% to 75% of the data will be located within a distance of one sigma unit on either side of the average; usually 90% to 98% of the data will be located with a distance of two sigma units on either side of the average; and approximately 99% to 100% of the data will be located within a distance of three sigma units on either side of the average. Thus, SPC methods have the advantage of being insensitive to distribution of the data of each of the subgroups because of the robust nature of using three-sigma limits about the mean.6,7,13 In addition, construction of the limits is less dependent on the measured process being in statistical control.13 Third, the conceptual foundation of control charts is rational sampling and rational subgrouping.13 Two conditions are required for any subgrouping to be rational: each subgroup must be logically homogeneous, and the variation within subgroups must be the logical and proper yardstick for setting limits on the variation between subgroups.14 Fourth, control charts are effective only to the extent that the organization can use, in an effective manner, the knowledge gained from the charts.13
The essence of SPC comprises simplicity and insight: the calculations require only a calculator, a table of correction factors for analysis,5-7,13 and graph paper for presentation.7 When data are collected in rational subgroups, three-sigma limits can be calculated and plotted for the subgroup averages, the subgroup dispersion statistic (range), and the individual data within all subgroups.6,7,13 An estimate of the dispersion parameter (sigma) can be made from any of at least seven statistics: average range, median range, average root mean square (RMS), median RMS, average SD, median SD, and pooled variance.13 After the dispersion statistic is calculated, it is multiplied by its appropriate correction factor to estimate three-sigma. For the same data set, any method for dispersion estimation will be sufficient. The average range requires the least rigor: subtract the lowest subgroup value from the highest subgroup value. The purpose of analysis is insight rather than numbers,13 as the objective is not "to compute the right limits" but rather "to take the right action upon the process"13; that is, to properly interpret the data from the process. Thus, the average range is used as an estimator of sigma.
The charts thus created are known as the average chart (for differences between the subgroups), the range chart (for checking consistency within the subgroups), and the individual-value histogram (which measures differences among the individual data values).13 When all data (group averages or ranges) fall within three-sigma limits, the process from which the data were obtained is said to be predictable.6,7,13 A process is defined as unpredictable when data obtained from the process occur in one of four patterns, known as signals.6
A signal is a single data point that falls outside the upper or lower three-sigma limits, or a series of data points that occur with a nonrandom pattern of variation around the central line. Signals were defined using the following rules6: Rule 1, one data point falls outside the upper or lower three-sigma limits; Rule 2, two of three consecutive data points fall on the same side of the average and beyond the two-sigma limits; Rule 3, four of five consecutive data points fall on the same side of the average and beyond the one-sigma limits; Rule 4, eight consecutive data points fall on the same side of the average. A process has undergone a detectable change if at least one of these situations exists.13
These criteria can also be used to determine if one process is different from another by comparing the data points of one process to the limits of the other. If the data points from an average or range value for one process show a signal when compared to the limits calculated for the other process, the two processes are said to be detectably different from each other.
In 1993, food-borne infections due to pork contaminated with Salmonella serovar Infantis prompted the Danish government to institute a Salmonella Combat Program.15 Features of this program included serologic testing of breeding and market swine.15 An indirect enzyme-linked immunosorbent assay (ELISA) was developed using lipopolysaccharide from Salmonella serovars Typhimurium and Choleraesuis (mix-ELISA).16 The mix-ELISA has been used to test serum and meat juice.17 Commercial swine herds are assigned to one of four Salmonella levels (categories) based on seroprevalence, ie, the proportion of samples per sampling with OD% > 20.18 The originally published cutoff value was OD% > 10.16
The purposes of this study were to confirm previously published results16 of studies on the mix-ELISA, to evaluate SPC methods for diagnostic test validation, and thus to demonstrate how serologic results could be used to evaluate Salmonella status of groups of pigs prior to slaughter.
Materials and methods
Animals
Forty 17-day-old pigs were purchased from one sow farm which had no history of clinical salmonellosis nor use of any Salmonella vaccine. Each pig was individually identified by a numbered ear tag and randomly assigned to one of four treatment groups (n = 10): one negative control group and three inoculated groups, each inoculated with one of three Salmonella serovars, Typhimurium, Choleraesuis, or Infantis. Each treatment group was randomly assigned to one of four individual isolation rooms, each containing one 5.5-m2 pen with solid flooring. Water was provided ad libitum and feed was provided daily.
This study was conducted in the Iowa State University Livestock Infectious Disease Isolation Facility. Experimental protocols were approved by the Iowa State University Committee on Animal Care.
Pre-inoculation sample collection
Blood samples and rectal swabs were collected from all pigs upon receipt, weekly for 9 weeks prior to inoculation, and on the day of inoculation (Day 0). On each sampling occasion, pooled fecal samples (five approximately 5-g samples) were collected from each pen floor. Rectal swabs and pooled floor feces were cultured for Salmonella. In order to confirm negative Salmonella status, a total of 24 pigs were randomly selected for euthanasia and necropsy prior to infection, one pig from each treatment group on Days -53, -46, -25, and -18, and two pigs from each treatment group on Day -4. We were unable to draw blood from a total of five different pigs: one pig on Day -63, three pigs on Day -60, and one pig on Day -53. Each pig was exsanguinated after euthanasia, and blood was collected for serological testing for Salmonella antibody.
Inoculation with Salmonella serovars
The strains of Salmonella Typhimurium, Infantis, and Choleraesuis var Kunzendorf that were used to inoculate the pigs were obtained during field studies from groups of market-weight pigs with no history of clinical salmonellosis.19 There was no history of Salmonella vaccine use on any of the farms from which the strains were obtained.
The remaining 16 pigs (four per treatment group) were approximately 77 days old when inoculated (Day 0). Feed was withheld from the inoculated pigs on Day 0. Each pig was manually restrained and 1.0 mL of inoculum (approximately 108 organisms) was sprayed into one nostril with a disposable syringe and Teflon cannula. The group inoculated with Salmonella Infantis was re-inoculated on Day 40 to attempt to produce reference sera for later studies.
Sampling post inoculation and termination of the experiment
Blood and rectal swabs were collected from all 16 pigs immediately prior to inoculation on Day 0. Pooled pen fecal samples were also collected on Day 0. After inoculation, blood and pooled pen fecal samples were collected on Days 3, 10, 17, 24, 31, 38, and 45; rectal swabs were collected on Days 0 through 5, 10, 17, 24, 31, 38, and 45.
The group inoculated with Salmonella Typhimurium were euthanized on Day 24 (in order to have reference sera from a time when Salmonella antibody was elevated) and the other three groups on Day 45. Pigs were exsanguinated at euthanasia and necropsies were performed. Samples of tonsil, lung, liver, spleen, jejunum, ileum, ileocecal lymph node, cecum, and colon were submitted for culture of Salmonella.
Blood and fecal sample testing
All serum samples were tested for Salmonella antibody in duplicate, over the course of 2 days, using the mix-ELISA.16 Results were reported as OD%, from which cutoff values were to be determined.
Samples were cultured for Salmonella as previously described.20 Presumptive Salmonella colonies were tested for "O" antigens by agglutination with serogroup antisera, and isolates were submitted for serotyping to the National Veterinary Services Laboratory (US Department of Agriculture, Ames, Iowa).
Preparation of SPC charts
Average and range charts were created for each treatment group according to Wheeler.6,7,13 The OD% values for each treatment group were subgrouped (n = 4) by date of blood sampling, and these data were used to assess the serologic status of treatment groups. Results from the negative control group were used to determine the cutoff for group average OD%, group OD% range, and individual OD% value. True Salmonella status of individuals was defined by culture results. A pig was considered true-positive if Salmonella was detectable either by rectal swab or organ culture. A pig was considered true-negative if Salmonella was not detectable either in a rectal swab or organ culture.
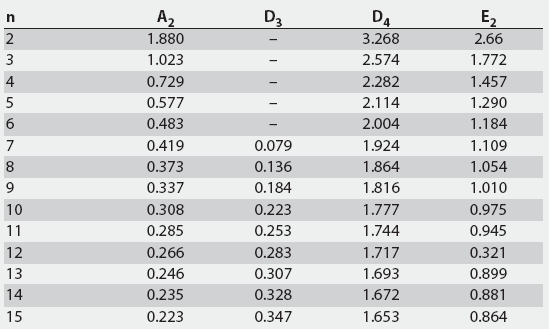
The upper control limits (UCL) and lower control limits (LCL) for the grouped data and the upper natural process limits (UNPL) and lower natural process limits (LNPL) for the individual data were calculated using three-sigma limits for group averages and ranges as well as individual values.13 The constants A2, D3, D4, and E2 are calculated from bias correction factors (Figure 1).6,7,13 For a subgroup of four, there is no D3 and the lower range limit (LRL) is 0.13
Figure 1: Calculation of control limits for average and range (X-bar - R) charts used to determine the cutoff values of the mix-ELISA in this study. The upper control limit (UCL) for the X-bar chart is calculated as UCL X-bar = X-bar + (A2 x R-bar) and the lower control limit (LCL) as LCL X-bar = X-bar - (A2 x R-bar), where X-bar = the grand average of the subgroup averages. The UCL and LCL for the R charts are calculated as UCLR = D4 x R-bar and LCLR = D3 x R-bar, respectively. The upper natural process limit (UNPL) for individual values is calculated as UNPLX = X-bar + (E2 x R-bar) and the lower natural process limit (LNPL) as LNPLX = X-bar - (E2 x R-bar), where X-bar = the average of the individual values. Factors for these formulas, given k subgroups each with n observations, are provided. For n greater than 15, refer to Wheeler, 1995.13 Reprinted with permission from SPC Press. Copyright 2004, SPC Press, Knoxville, Tennessee. All rights reserved.
|
For the average OD% of each treatment group,
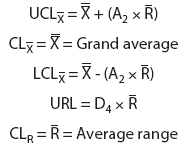
For individual values within the negative control group,
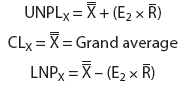
After these experiments, infected and noninfected processes were defined. The infected process was defined as pigs infected with Salmonella (inoculated and confirmed infected by culture). The noninfected process was defined as pigs not infected with Salmonella (not inoculated and confirmed not infected by culture).
The UCL, upper range limit (URL), and UNPL for the negative controls were used as the basis for cutoff determination. A conservative approach was used to calculate the cutoff by approximating the one-sided 95% confidence interval of the respective upper limits. Thus, the group average OD% cutoff was calculated as follows:

where Zone-sided0.95 = 1.645; sd = the standard deviation of the 24 OD% values for the negative control pigs from Days 3, 10, 17, 31, 38, and 45; and N = 24, the number of all OD% values in the negative control group.
The group range cutoff was calculated as follows from the range values of the negative control group:

The cutoff for individual values was calculated as follows:

Serologic sensitivity and specificity were calculated by considering the one-sided 95% confidence interval for the UNPL of individual serologic values and the known culture status of the negative control group. As OD% values were reported as whole numbers,16 a calculated UNPL plus the one-sided 95% confidence interval that was not a whole number was rounded down for use as the cutoff. A treatment group was determined to be seropositive if one pig's OD% was greater than this cutoff. Seroprevalence for each treatment group was calculated as a proportion of the number of seropositive pigs in the group, using the UNPL for individual values as the cutoff.
ROC analysis was performed on the individual OD% results for each postinoculation day and on all individual results in the experiment, using SPSS Base Version 7.5 (SPSS Inc, Chicago, Illinois).
Results
Bacteriology
All individual pig rectal swabs and pooled fecal samples from the pen floors that were collected prior to inoculation, and all organ samples collected from pigs euthanized prior to Day 0, were culture-negative for Salmonella.
All pooled fecal samples (pen feces) from the Salmonella Choleraesuis group and from the negative control pigs, from Day 0 through the end of the study, were culture-negative for Salmonella. In the Salmonella Typhimurium group, Salmonella Typhimurium was detected in pen feces only on Day 23, and in the Salmonella Infantis group, Salmonella Infantis was detected in pen feces on Days 17, 23, and 38.
Salmonella was isolated from at least one rectal swab in all three groups of inoculated pigs, and in each case, the isolates were the homologous serotypes. All pigs inoculated with either Salmonella Typhimurium or Salmonella Infantis shed the homologous serotype. In the Salmonella Typhimurium group, the rectal swab from one pig was culture-positive for Salmonella Typhimurium on Day 23. In the Salmonella Choleraesuis group, rectal swabs from one pig on Day 1 and another pig on Days 3 and 4 were culture-positive for Salmonella Choleraesuis. In the Salmonella Infantis group (re-inoculated on Day 40), rectal swabs from all four pigs were culture-positive for 4 days after infection, then at least once after re-inoculation.
All organ samples collected from the negative control pigs were culture-negative for Salmonella. In the Salmonella Typhimurium and Salmonella Infantis groups, at least one organ of all four pigs was culture-positive for the homologous serotype. In the Salmonella Choleraesuis group, at least one organ was culture-positive in only two of the four pigs.
Serological results
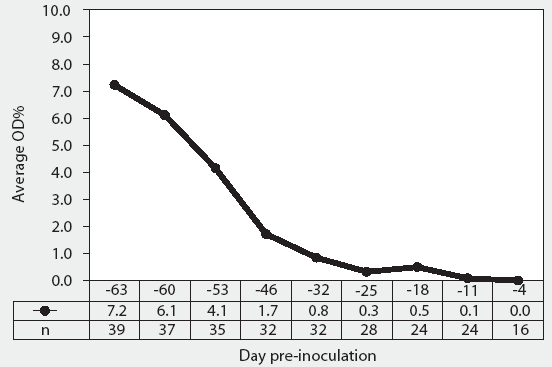
Figure 2 summarizes the time course of Salmonella antibody production detected by the mix-ELISA (expressed as average OD%) in all pigs sampled prior to inoculation. In some pigs, OD% was detectable at Day -63. Average OD% declined during the course of the pre-infection period (Days -63 to 0).
Figure 2: Blood samples were collected weekly from forty 17-day-old pigs (culture-negative for Salmonella) upon receipt, for 9 weeks prior to inoculation with Salmonella serovars, and on the day of inoculation at approximately 77 days of age (Day 0). Pigs were divided into four equal treatment groups housed in separate rooms. A total of 24 pigs were euthanized prior to inoculation: one pig from each treatment group on Days -53, -46, -25, and -18, and two pigs from each treatment group on Day -4. We were unable to draw blood from a total of five different pigs: one pig on Day -63, three pigs on Day -60, and one pig on Day -53. Serum samples were tested for Salmonella antibody using the mix-ELISA and results were reported as OD%. Average OD% is the average for all animals tested on the day of sampling (n).
|
Figures 3 through 6 summarize the postinoculation time course of antibody production for all four treatment groups (n = 4) plotted on average and range charts. The OD% increased as early as 3 days post inoculation in at least one pig in each group inoculated with Salmonella.
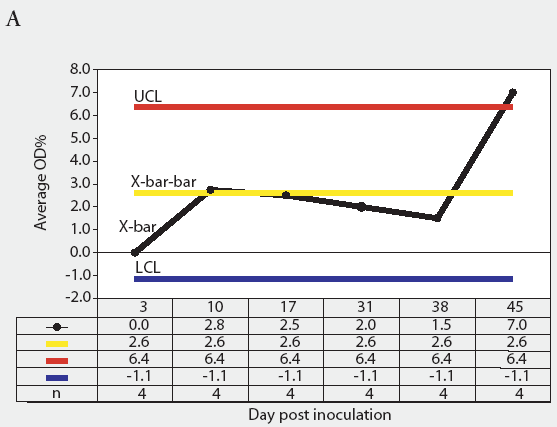
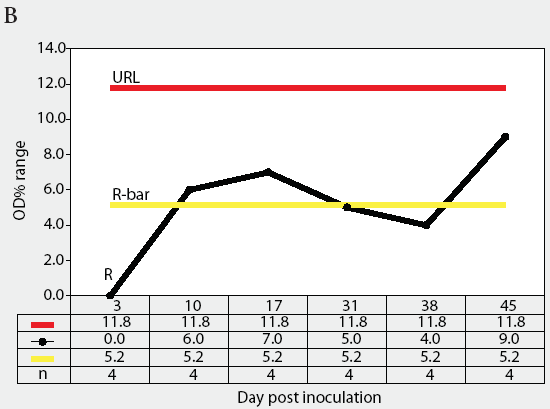
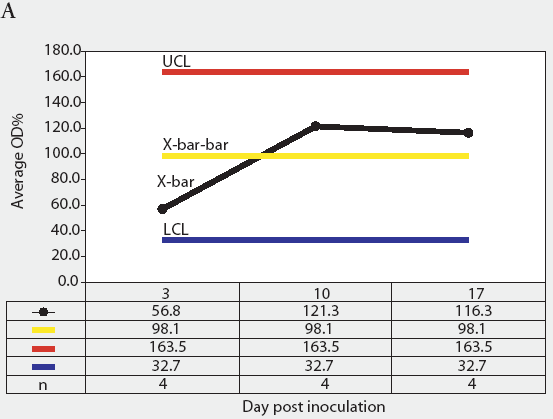
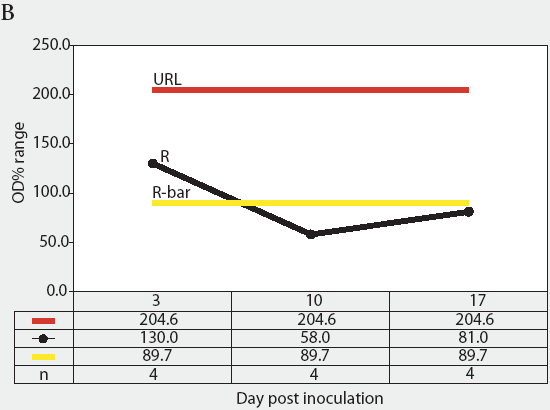
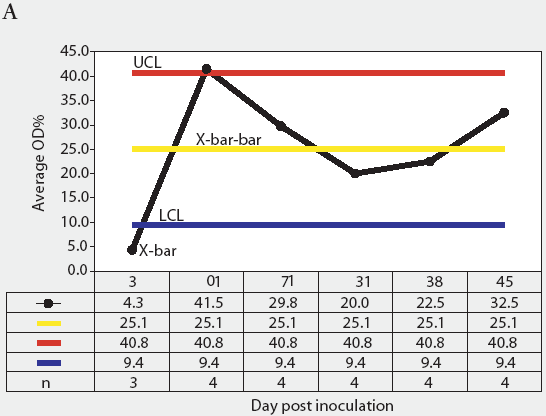
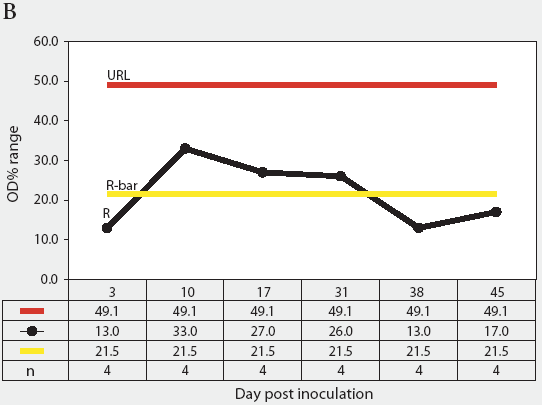
Figure 3: Forty 17-day-old pigs (culture-negative for Salmonella) were evenly assigned to four treatment groups housed in separate rooms and inoculated with Salmonella serovars Typhimurium, Infantis, or Choleraesuis var Kunzendorf, or not inoculated (negative control group), at approximately 77 days of age (Day 0). Blood samples were collected upon receipt, weekly for 9 weeks prior to inoculation with Salmonella (Day 0), and weekly until termination of the experiment, which was Day 45 for all groups except the serovar Typhimurium group, which was euthanized on Day 24. Serum samples were tested for Salmonella antibody using the mix-ELISA. Results were reported as OD% and subgrouped by date of collection. Statistical process control charts of mix-ELISA results from the negative controls are shown: Figure 3A, X-bar chart of OD% values and Figure 3B, range chart of OD% values. X-bar is the subgroup average; X-bar-bar is the grand average of the subgroups; UCL and LCL are the upper and lower control limits, respectively, of the X-bar chart; n is the number of animals per subgroup; R is the range of data for each subgroup; R-bar is the average range for all subgroups; and URL is the upper range limit for the R chart.
|
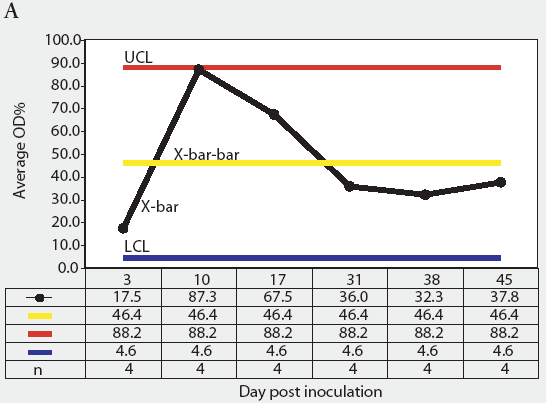
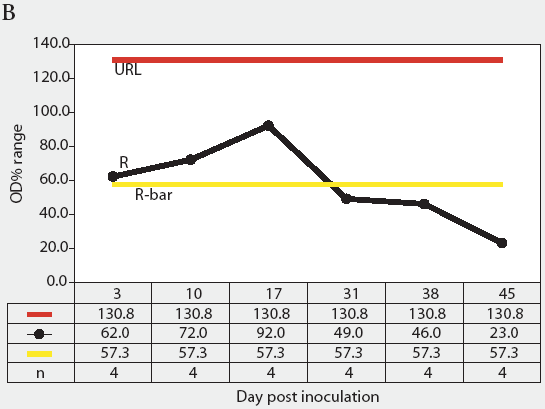
Figure 4: Statistical process control charts of the mix-ELISA results from the Salmonella Typhimurium-infected group of pigs in the study described in Figure 3: Figure 4A, X-bar chart of OD% values and Figure 4B, range chart of OD% values. X-bar is the subgroup average; X-bar-bar is the grand average of the subgroups; UCL and LCL are the upper and lower control limits, respectively, of the X-bar chart; n is the number of animals per subgroup; R is the range of data for each subgroup (subgrouped by date of blood collection); R-bar is the average range for all subgroups; and URL is the upper range limit for the R chart.
|
Figure 5: Statistical process control charts of the mix-ELISA results from the Salmonella Choleraesuis-infected group of pigs in the study described in Figure 3: Figure 5A, X-bar chart of OD% values and Figure 5B, range chart of OD% values. X-bar is the subgroup average; X-bar-bar is the grand average of the subgroups; UCL and LCL are the upper and lower control limits, respectively, of the X-bar chart; n is the number of animals per subgroup; R is the range of data for each subgroup (subgrouped by date of blood collection); R-bar is the average range for all subgroups; and URL is the upper range limit for the R chart.
|
Figure 6: Statistical process control charts of the mix-ELISA results from the Salmonella Infantis-infected group of pigs in the study described in Figure 3: Figure 6A, X-bar chart of OD% values and Figure 6B, range chart of OD% values. X-bar is the subgroup average; X-bar-bar is the grand average of the subgroups; UCL and LCL are the upper and lower control limits, respectively, of the X-bar chart; n is the number of animals per subgroup; R is the range of data for each subgroup (subgrouped by date of blood collection); R-bar is the average range for all subgroups; and URL is the upper range limit for the R chart.
|
The OD% values from the negative controls were used to determine three serologic cutoffs: group average OD% = 7.239, group range OD% = 12.612, and individual OD% = 11.024. As OD% results are reported as whole numbers, the individual OD% cutoff was 12.
When the definition of true Salmonella status was applied, one pig inoculated with Salmonella Infantis that was never culture-positive was excluded from the calculation of sensitivity and specificity for individual values. Mix-ELISA sensitivity was 0.40 (95% CI, 0.046 - 0.754) on Day 3; 1.00 (95% CI, 0.75 - 1.00) on Day 10; 0.86 (95% CI, 0.455 - 1.00) on Day 38; and 1.00 (95% CI, 0.75 - 1.00) on Day 45. Mix-ELISA specificity was 1.00 (95% CI, 0.75 - 1.00) for the same 4 days. When individual results from all postinoculation days were combined, sensitivity was 0.85 (95% CI, 0.36 - 1.00) and specificity was 1.00 (95% CI, 0.96 - 1.00).
When seroprevalence was the criterion to assess group Salmonella status, all inoculated groups were correctly identified as positive (seroprevalence > 0) and all noninoculated groups were correctly identified as negative (seroprevalence = 0).
The average OD% and OD% range of the three inoculated groups for each postinoculation day, with the exception of the average OD% of the Salmonella Infantis group at Day 3, were greater than the UCL (7.239) on the SPC chart of the negative control pigs from Day 3 until the end of the study. These are Rule 1 signals, interpreted as meaning that the OD% averages from these days were not characteristic of Salmonella-negative pigs, and the pigs were therefore considered exposed to Salmonella. The OD% of the Salmonella Infantis group on Day 3 (4.3) was less than 7.239, but the OD% range of this group (13) was greater than that of the negative control pigs for that day (12.617). Thus, this group was correctly identified as exposed to Salmonella. The negative controls were correctly identified as Salmonella-negative on all postinoculation days.
When the SPC charts for the Salmonella Infantis and Salmonella Choleraesuis groups were compared to those of the Salmonella Typhimurium group, the Salmonella Choleraesuis-infected group showed a Rule 1 signal on Day 38, Rule 2 signals on Days 17, 31, 38, and 45, and Rule 3 signals on Days 3, 10, 17, 31, 38, and 45. The Salmonella Infantis group showed Rule 1 signals on Days 17, 31, 38, and 45, and Rule 2 and 3 signals on Days 3, 10, 17, 31, 38, and 45. Thus, the serologic responses of the Salmonella Infantis and Salmonella Choleraesuis groups differed from that of the Salmonella Typhimurium group.
The areas under the ROC curve for individual OD% values on each postinoculation day were 0.286 for Day 3, 0.00 for Day 10, 0.00 for Day 38, and 0.00 for Day 45. The area under the ROC curve for all individual OD% values was 0.936 (95% CI, 0.878 - 0.994), with sensitivity optimised at 0.849 and specificity optimised at 1.00 when the cutoff was OD% >= 11.5. When the cutoff was OD% >= 12.5, sensitivity was 0.830 and specificity remained at 1.00.
Discussion
This study confirmed that the mix-ELISA detects antibody from pigs infected with Salmonella16,21in a laboratory setting. The SPC methods used in this study were able to detect infected groups 3 days after infection. Nielsen et al (1995)16 detected seroconversion in feeder pigs 7 days post inoculation and Harris21 detected seroconversion 8 days post inoculation. Thus, we would expect the mix-ELISA to detect populations of swine exposed to Salmonella serovars within the last 3 to 8 days of finishing. The evidence of this exposure in a swine population will be one of the following criteria: average OD% greater than the upper 95% confidence interval of the UCL of average OD% from a known-negative population; range of OD% greater than the upper 95% confidence interval of the UCL average range values for a known-negative population; or prevalence > 0, determined by the UNPL for individuals from a known-negative population. Additional testing of known-negative populations is recommended to determine the cutoff and assay characteristics determined herein.12
Statistical process control methods are rational and objective for determining the cutoffs for diagnostic assays. Data from known-negative populations serve as the foundation for constructing the cutoff. When these data are collected in rational subgroups, k, containing n observations per subgroup, control limits are calculated from approximately 0.9k (n - 1) degrees of freedom.13 When the degrees of freedom are >10, the calculated control limits are "set" and are not appreciably changed by additional degrees of freedom.13 The limits for this study were calculated from 16.2 degrees of freedom. An upper 95% confidence interval can be calculated to add a conservative factor for a cutoff, thus reducing the likelihood of making Type I errors (false-positive) in data interpretation.
Because rational subgrouping is used to calculate SPC limits (and is foundational for calculating SPC limits), one may then make objective conclusions about the populations from which the samples were collected. An investigator evaluates the location statistic (eg, average OD%) and the dispersion statistic (eg, OD% range) to determine the status of each sampled population compared to its respective limits. Since the cutoff is determined by testing known-negative swine, thus characterizing true-negative populations and true-negative individuals, interpretation of the results can be made without regard for the time when pigs were exposed to Salmonella. Group average or range values or individual values that lie outside these limits are not characteristic of negative groups or individuals, and thus the interpretation is that these pigs are exposed. Therefore, SPC methods provide both a herd-level and individual-level method for evaluating diagnostic assay results.
Statistical process control methods clearly provide three different cutoffs for evaluating data: subgroup average OD%, subgroup OD% range, and individual OD%. While other methods are recognized for determining cutoff,9,11 SPC methods have the advantage of being simple and insensitive to data distribution. For instance, ROC curve analysis can be used to determine an optimized cutoff. In this study, ROC curve analysis provided the same cutoff as did SPC methods when all individual data were analysed. ROC analysis required a statistical software program, did not take into account the group average or dispersion (range) statistic as parameters to assess group status, and did not analyse each subgroup. The SPC values can be determined with pencil, paper, and a table of average and range chart factors. The robust nature of SPC methods stems from the use of three-sigma limits, making the distribution of subgroup data irrelevant to the accuracy and utility of calculated limits.13 Thus, data transformation to normalize data is also not necessary.13
When SPC methods are used, a Salmonella control-and-reduction program can be introduced without concern for capricious minimum levels of exposure. Rather than arbitrarily deciding to set a minimum level of exposure to Salmonella, production systems can compare their serologic data to known-negative populations of market pigs. Then they can continue to monitor Salmonella serostatus and measure the effectiveness of interventions by first describing the current level of exposure to Salmonella. Thus, a system has created the three components of an operational definition22 to evaluate its Salmonella control and reduction. An operational definition22 has three components: a stated objective (reduction and control of Salmonella); a method of measuring progress toward that objective (ongoing serological testing using the mix-ELISA); and a method to assess whether the objective has been or is being met (SPC charts and the use of the interpretation rules).22
We clearly showed that the immune response produced after exposure to Salmonella Typhimurium was different from that elicited by exposure to Salmonella Choleraesuis and Salmonella Infantis. The implication of this in a Salmonella reduction program is that a reduction in OD% might be indicative of a change in the serotype present in the herd rather than an effect associated with an intervention strategy. One would expect that if a new serotype were introduced, herd SPC charts would remain constant in the absence of additional intervention strategies. In the presence of an intervention strategy, the farm management team would need to determine if a change in serotype had occurred, then re-evaluate and redesign a strategy for further decreasing Salmonella exposure and measure that reduction via a reduction in OD%. In this study, the mix-ELISA results of the pre-inoculation serum samples indicate that there were some seropositive pigs received for this experiment. Since there was no bacteriological evidence of Salmonella from any rectal swab or pooled pen samples, we conclude that these pigs either suckled seropositive sows or had been infected with Salmonella and ceased shedding prior to receipt.
An objection to use of serological testing to assess Salmonella status is that a pig can become infected and shed in as little as 4 hours post infection,23 implying that low-level exposure swine could be infected in lairage when brought into contact with high-level exposure swine. Serologic monitoring of populations of swine over time using SPC methods would enable production systems and abattoirs to identify either low-level or high-level exposure sources of pigs and to make decisions that might prevent cross-contamination.
Statistical process control methods might also be used to assess the manufacturing process of any serologic assay kit or components, thus permitting predictable results between and within laboratories. When a diagnostic-kit manufacturing company can display statistical control (absence of signals) in its processes for assay manufacturing, one would expect that end-users would be able to predictably perform the assay. In the absence of predictable results from a laboratory, SPC methods provide an objective basis for investigating possible causes of the lack of predictability. Once the causes have been removed, and the assay results become predictable, results can be confidently reported by the laboratory. In this scenario, it would be expected that the following sources of variation could be controlled: serial-to-serial; laboratory-to-laboratory; technician-to-technician; and day-to-day. Studies are in progress to test this system of consistent (predictable) assay manufacturing processes and consistent (predictable) results across the above sources of potential variation.
Implications
- () Statistical process control methods can be used to evaluate the characteristics of the Salmonella mix-ELISA.
- () Statistical process control methods and the mix-ELISA can be used as tools for a Salmonella monitoring strategy.
Acknowledgements
The authors thank Ellen Martens who completed all of the serologic testing for this study, Kelly Burkhart who completed all of the bacteriologic culture work for this study, and David Brown for the care of the animals during the course of this study. This study was supported in part by the United States Department of Agriculture Fund for Rural America and the Iowa Agriculture Experimental Station.
References
1. Kuby J. Immunology. New York: WH Freeman and Company; 1999.
2. Tyler JW, Cullor JS. Titers, tests, and truisms: rational interpretation of diagnostic serologic testing. JAVMA. 1989;194:1550-1558.
3. Shewhart WA. Economic Control of Quality of Manufactured Product. New York, New York: D. Van Nostrand Inc; 1931:501.
4. Shewhart WA. Statistical Method from the Viewpoint of Quality Control. Mineola, New York: Dover Publications Inc; 1939.
5. Wheeler DJ. Understanding Industrial Experimentation. 2nd ed. Knoxville, Tennessee: SPC Press Inc; 1990.
6. Wheeler DJ. Understanding Statistical Process Control. Knoxville, Tennessee: SPC Press Inc; 1992.
7. Wheeler DJ. Understanding Variation. The Key to Managing Chaos. Knoxville, Tennessee: SPC Press Inc; 1993.
8. Martin SW, Meek AH, Willeberg P. Veterinary Epidemiology: Principles and Methods. Ames, Iowa: Iowa State University Press; 1987.
9. Smith RD. Veterinary Clinical Epidemiology: A Problem-Oriented Approach. 2nd ed. Boca Raton, Florida: CRC Press Inc; 1995.
10. Thrushfield M. Veterinary Epidemiology. 2nd ed. Oxford, UK: Blackwell Science; 1995.
11. Greiner M, Pfeiffer D, Smith RD. Principles and practical application of receiver-operating characteristic analysis for diagnostic tests. Prev Vet Med. 2000;45:23-41.
12. Jacobson RH. Validation of serological assays for diagnosis of infectious diseases. Rev Sci Tech Off Int. 1998;17:469-486.
13. Wheeler DJ. Advanced Topics in Statistical Process Control. The Power of Shewhart's Charts. Knoxville, Tennessee: SPC Press Inc; 1995.
14. Wheeler DJ. Classics from Donald J. Wheeler. The Process Behaviour Chart Papers. Knoxville, Tennessee: SPC Press Inc; 1999.
15. Mousing J, Jensen PT, Halgaard C, Bager F, Feld N, Nielsen B, Nielsen JP, Bech-Neilsen S. Nation-wide Salmonella enterica surveillance and control in Danish slaughter swine herds. Prev Vet Med. 1997;29:244-261.
16. Nielsen B, Baggesen D, Bager F, Haugegaard J, Lind P. The serological response to Salmonella serovars typhimurium and infantis in experimentally infected pigs. The time course followed with an indirect anti-LPS ELISA and bacteriological examinations. Vet Microbiol. 1995;47:205-218.
17. Nielsen B, Ekeroth L, Bager F, Lind P. Use of muscle juice as a source of antibodies for serological detection of Salmonella infection in slaughter pig herds. J Vet Diagn Invest. 1998;10:158-164.
*18. Ekeroth L, Alban L, Feld N. Single testing versus double testing of meat juice samples for Salmonella. Proc IPVS. Ames, Iowa. 2002;1:312.
19. Baum DH. Vaccine and epidemiological studies of Salmonella in swine in microbiology, immunology, and preventive medicine [PhD dissertation]. Ames Iowa: Iowa State University; 1997.
20. Roof MB, Doitchinoff DD. Safety, efficacy, and duration of immunity induced in swine by use of an avirulent live Salmonella choleraesuis-containing vaccine. AJVR. 1995;56:39-44.
*21. Harris IT. Serological basis for assessment of subclinical Salmonella infection in swine: Part I. J Swine Health Prod. 2003;11:247-252.
22. Wheeler DJ, Poling S. Building Continuous Improvement. Knoxville, Tennessee: SPC Press Inc; 1998.
23. Fedorka-Cray PJ, Kelly LC, Stable TJ, Gray JT, Laufer JA. Alternate routes of invasion may affect pathogenesis of Salmonella typhimurium. Inf Imm. 1995;63:2658-2664.
* Non-refereed references.