Original research |
Peer reviewed |
Porcine reproductive and respiratory syndrome virus: Age and management system disease modeling for pathogenic co-infection
Virus del síndrome reproductivo y respiratorio porcino: Modelo de un sistema de enfermedad por edad y manejo de sistema para una coinfección patogénica
Virus du syndrome reproducteur et respiratoire porcin: Étude de modélisation selon l’âge et les systèmes de régie sur les co-infections
Paul M. Dorr, DVM, PhD; Wondwossen A. Gebreyes, DVM, PhD, Diplomate ACVPM; Glen W. Almond, DVM, MSc, PhD
PMD, GWA: Department of Population Health and Pathobiology, College of Veterinary Medicine, North Carolina State University, Raleigh, North Carolina. WAG: Department of Veterinary Preventative Medicine, College of Veterinary Medicine, The Ohio State University, Columbus, Ohio. Corresponding author: Dr Paul M. Dorr, Merial Ltd, 6498 Jade Rd, Fulton, MO 65251; Tel: 573-642-5977, Ext 1109; Fax: 573-642-0356; E-mail: paul.dorr@merial.com. Dr Dorr was supported by internal funding, private agricultural industry, and Pfizer Animal Health.
Cite as: Dorr PM, Gebreyes WA, Almond GW. Porcine reproductive and respiratory syndrome virus: Age and management system disease modeling for pathogenic co-infection. J Swine Health Prod. 2007;15(5):258–263.
Also available as a PDF.
SummaryObjectives: To construct models evaluating statistically significant associations between porcine reproductive and respiratory syndrome virus (PRRSV) and co-infectious agents, and to assess effects of changes in age and management system on co-infection status, serological profiles, lung lesions, and histopathological lesions in PRRSV-positive pigs Materials and methods: Pigs 3, 9, 16, and 24 weeks of age from 41 sow sources representing one-, two-, and three-site production systems were necropsied on the basis of acute, subacute, and chronic disease stages (n = 5 animals per age group per farm). Logistic regression models were used to assess co-infection with other pathogens, serological profiles, and histopathological lesions in pigs PRRSV-positive by polymerase chain reaction and with lung lesions consistent with PRRSV infection, and in PRRSV-negative pigs. Additional models were stratified by age and type of production system. Results: In the overall model, clinically ill PRRSV-positive pigs were more likely than PRRSV-negative pigs to be co-infected with swine influenza virus (SIV) type A (OR, 2.40; 95% CI, 1.07-5.39) and to have lung scores in the 11% to 50% range (OR, 1.65; 95% CI, 1.06-2.57). Three-week-old PRRSV-positive pigs were more likely to be co-infected with porcine circovirus type 2 (OR, 8.83; 95% CI, 1.76-44.26). Nine- and 16-week-old pigs were 15.57 and 5.75 times as likely to be co-infected with SIV, respectively. Implications: Changes in PRRSV infection, disease manifestation, and associated co-factors such as age and type of management system are important considerations for prevention, vaccination timing, diagnostic testing, and treatment for disease complexes associated with PRRSV. | ResumenObjetivos: Construir modelos que evalúen asociaciones estadísticamente significativas entre el virus del síndrome reproductivo y respiratorio (PRRSV por sus siglas en inglés) y agentes coinfecciosos, y evaluar los efectos del cambio de edad y sistema de manejo en el estatus de coinfección, perfiles serológicos, lesiones pulmonares y lesiones histopatológicas en cerdos positivos al PRRSV. Materiales y métodos: Se realizó una necropsia a cerdos
de 3, 9, 16, y 24 semanas de edad de 41 fuentes de hembras que representaban
a sistemas de producción de uno, dos, y tres sitios en base a los estados
de enfermedad aguda, subaguda, y crónica Resultados: En el modelo total, los cerdos clínicamente enfermos y positivos al PRRSV fueron más propensos que los cerdos negativos al PRRSV a ser coinfectados con el virus de la influenza porcina (SIV por sus siglas en inglés) tipo A (OR, 2.40; 95% CI, 1.07-5.39) y a tener porcentajes pulmonares en el rango de 11% a 50% (OR, 1.65; 95% CI, 1.06-2.57). Los cerdos positivos al PRRSV de 3 semanas de edad fueron más propensos a ser coinfectados con el circovirus porcino tipo 2 (OR, 8.83; 95% CI, 1.76-44.26). Los cerdos de 9 y 16 semanas de edad fueron 15.57 y 5.75 veces más propensos a ser coinfectados con el SIV, respectivamente. Implicaciones: Los cambios en la infección con el PRRSV, manifestaciones de la enfermedad, y factores coasociados tales como el tipo de sistema de manejo y edad son consideraciones importantes para la prevención, tiempo de vacunación, prueba de diagnóstico, y tratamiento para los complejos de la enfermedad asociados con el PRRSV. | ResuméObjectifs: Élaborer des modèles évaluant les associations statistiquement significatives entre le virus du syndrome reproducteur et respiratoire porcin (PRRSV) et des agents de co-infections, et déterminer les effets de changements dans l’âge des animaux et des systèmes de régie sur le statut des co-infections, les profils sérologiques, les lésions pulmonaires, et les lésions histopathologiques chez des porcs positifs pour le PRRSV. Matériels et méthodes: Des porcs âgés de
3, 9, 16, et 24 semaines provenant de 41 truies représentant des systèmes
de production sur un, deux, et trois sites ont été soumis à une
nécropsie sur la base du stade de la maladie (aigu Résultats: Dans le modèle général, les porcs PRRSV-positifs et cliniquement malades étaient plus susceptibles que les porcs PRRSV-négatifs à être co-infectés par le virus de l’influenza porcin (SIV) de type A (OR, 2.40; IC 95%, 1.07-5.39) et d’avoir des pointages de lésions pulmonaires dans la plage de 11% à 50% (OR, 1.65; IC 95%, 1.06-2.57). Les porcs PRRSV-positifs âgés de 3 semaines étaient plus susceptibles d’être co-infectés avec le circovirus porcin de type 2 (OR, 8.83; IC 95%, 1.76-44.26). Une co-infection avec le SIV était 15.57 et 5.75 fois plus probable de se produire chez les porcs âgés respectivement de 9 et 16 semaines. Implications: Des modifications dans l’infection par le PRRSV, les manifestations de la maladie, et des cofacteurs associés tels que l’âge et le type de système de régie sont des considérations importantes pour la prévention, le moment de la vaccination, les épreuves diagnostiques, et le traitement de complexes de maladies associés au PRRSV.
|
Keywords: swine, porcine
reproductive and respiratory syndrome virus, PRRSV, co-infection, disease
modeling, epidemiology
Search the AASV web site
for pages with similar keywords.
Received: February
20, 2007
Accepted: May
22, 2007
Since its discovery in 1987,1 porcine reproductive and respiratory syndrome virus (PRRSV) has caused major health, management, and financial burdens in the swine industry. In the United States, a 2005 study by Neumann et al2 estimated that PRRSV is responsible for approximately $560 million in producer losses annually. Even though it is considered a primary pathogen, PRRSV rarely acts alone in growing pigs. It is one of several major contributors to the porcine respiratory disease complex (PRDC).3 The emergence of PRRSV, in addition to the change in pathogenicity of porcine circovirus type 2 (PCV2) over the past 5 to 6 years, coupled with the pathologic associations of these two viruses,4 may have significantly increased PRRSV costs since the Neumann et al study2 was completed.
Many bacterial and viral pathogens may co-infect pigs with porcine reproductive and respiratory syndrome (PRRS) in clinical and experimental settings. Co-infectors include swine influenza virus (SIV), PCV2, porcine respiratory coronavirus, Mycoplasma hyopneumoniae, Salmonella serovar Choleraesuis, Haemophilus parasuis, Streptococcus suis, Actinobacillus pleuropneumoniae, Bordetella bronchiseptica, and Pasteurella multocida. Some studies5-13 have shown that co-infection may potentiate respiratory disease or worsen the clinical outcome, while others suggest that there is no effect, or even a slight protective effect in some cases.14-16 The clinical expression of co-infectious agents varies with production system. These variables include, but are not limited to, the age of the animal, type of management system, infectious dose, reproductive rate of the virus within the herd, pathogen virulence, immune status of the herd, and biosecurity practices.3,4,12
The purpose of this study was to identify co-infectious agents in PRRSV-infected pigs and to evaluate significant associations between PRRSV and these agents by constructing disease models, assessing effects of changes in age and type of management system on co-infection status, serological profiles of co-infectors, and gross lung involvement and histopathological lesions of PRRSV-positive pigs.
Materials and methods
Animals and housing
Data for the disease models were derived from a cohort of pigs as previously reported.4 The study animals were obtained from a single integrated pig productionsystem subset of production flows (with production flow defined as a single-sow source and its associated nurseries and finishers in one-, two-, or three-site management systems). Study animals were selected from production flows in which nursery or finisher mortality or both approached or exceeded 10%, either on a recent or long-term basis. All production flows utilized conventional confinement housing systems. These data were originally used for clinical diagnostic purposes on individual farms within the production system.
Study design
This cross-sectional study included a group of 791 pigs, representing 41 production flows, over a period of approximately 3 months. Within each production flow, animals were selected for necropsy from four different age groups: 3-week-old pigs (early nursery), 9-week-old pigs (late nursery), 16-week-old pigs (early to mid-finishing), and 24-week-old pigs (late finishing). Five animals of each age group were selected for necropsy from a pool of animals within a single barn. Animals were selected by stage of disease (respiratory or enteric disease or both) and were assigned a disease score from 1 to 5 as follows: scores 1 and 2, well-fleshed animals in the peracute stages of the disease process; scores 3 and 4, animals in the acute to sub-acute stage of the disease process, showing clinical signs, and having poorer body condition than their penmates; score 5, chronically ill animals with considerable loss of body condition, but not located in a treatment or hospital pen. All pigs were humanely euthanized using a lethal intravenous barbiturate injection or captive bolt. The study protocol followed the guidelines of the North Carolina State University Institutional Animal Care and Use Committee.
Necropsy and diagnostics
All necropsies for each farm were completed on site, beginning with the youngest and moving to the older groups of pigs. Within each age group, pigs with disease score 1 were necropsied first, followed by pigs with scores 2, 3, 4, and 5. Separate instruments were used for enteric and non-enteric tissues, and were cleaned between necropsies. To assign a lung score, the affected proportion of each of the seven lung lobes was assessed. For each lobe, the percent affected was multiplied by the estimated lung capacity of that lobe (10% each for right and left cranial, right and left middle, and accessory lobes; 25% each for the right and left caudal lobes), and results for the seven lobes were added to obtain the total percent of lung affected. Lung scores were later categorized for modeling as low (0% to 10%), medium (11% to 50%), and high (51% to 100%). Fresh and fixed tissues were collected from each animal. Enteric and non-enteric tissues were stored in separate Whirl-Pak bags (Nasco, Fort Atkinson, Wisconsin), and a fresh liver sample was also stored in a separate bag. Fixed and fresh tissue samples included tonsil, hilar lymph nodes, lung, heart, spleen, kidney, liver, brain (3-week-old pigs), mesenteric lymph nodes, ileum, jejunum, cecum, and colon. A vial of whole blood was collected from each pig. All samples were submitted to the Veterinary Diagnostic Laboratory at the University of Minnesota (St Paul, Minnesota). Diagnostic methods for each case included polymerase chain reaction (PCR) for PRRSV (US and European), PCV2, SIV type A, and M hyopneumoniae. Electron microscopy was performed on fecal samples for porcine rotavirus. Immunohistochemical testing was performed on intestinal tissues for detection of transmissible gastroenteritis virus. Histological testing was performed on all tissues, and results were allotted into five categories, which included PRRSV lung lesions, M hyopneumoniae lung lesions, SIV lung lesions, small intestinal villous inflammation or atrophy-blunting, and “other”. The “other” category included non-enteric or non-lung lesions, eg, endocarditis, pericarditis, polyserositis, and hepatic, splenic, central nervous system, and kidney lesions. All fresh tissues were cultured for bacteria, and isolates were identified and tested for antimicrobial sensitivity. Serologic tests were performed on each blood sample for PRRSV, M hyopneumoniae, SIV H1N1, and SIV H3N2, using ELISAs for all pathogens except SIV H3N2, for which a hemagglutination inhibition (HI) test was used. Results of the ELISA and HI tests were entered into low, medium, and high categories for model inclusion. Categories for ELISA sample:positive (S:P) ratios included ranges of 0.00 to 0.49 (low, coded as 0), 0.50 to 0.99 (medium, coded as 1), and ≥ 1.00 (high, coded as 2). Categories of HI titers included ranges of ≤ 1:40 (low, coded as 0), 1:80 to 1:160 (medium, coded as 1), and ≥ 1:320 (high, coded as 2). Intestinal content was screened for parasites by direct examination and fecal flotation.
Statistical analysis
Diagnostic reports for 791 pigs were entered into a database. From this pool of pigs, data from 583 animals were included in the models. The remaining 208 animals were excluded either because of variation in age (not 3, 9, 16, or 24 weeks of age, n = 83) or because of missing lung scores or serological data (n = 125). For all animals included in the final analysis, data was complete for all variables evaluated, ie, 28 disease variables and three production variables (age, production flow, and disease score). Disease variables included results from PCR testing for specific pathogens, virus isolation, and bacterial culture; results from histological evaluations in five categories; lung lesions suggestive of infection with PRRSV, SIV, or M hyopneumoniae; enteric lesions; lesions categorized as “other” (included histopathological lesions suggestive of systemic infection, eg, peritonitis, pericarditis, polyserositis, hepatic and splenic lesions, and joint effusions); lung scores; and results of serological testing for PRRSV, SIV H1N1, SIV H3N2, and M hyopneumoniae. Animals were coded as PRRSV-positive if they met the following two criteria: PCR-positive for PRRSV, and positive for histopathological lesions suggestive of PRRSV infection. Animals meeting these criteria were compared to animals not meeting these criteria for statistically significant associations of co-infections, lesions, and serological changes. All statistical analyses were performed using EGRET statistical software (Cytel Inc, Cambridge, Massachusetts). Variables were initially screened using bivariate analysis. Traditional 2 � 2 tables on the 28 disease variables were constructed with the “outcome” variable defined as positive for PRRSV both by PCR and by observation of histopathological lesions suggestive of PRRSV infection. Exposure variables with �‡² P values ≤ .15 were retained for model fitting. Model fitting was performed using logistic regression by backwards elimination of all retained variables from the bivariate analysis. Next, a simple effects model was constructed. All variables were then tested for potential interactions. Potential confounders (age, type of production flow, and disease score) were analyzed in the overall model and subsequently in stratified models, and were included in models for which they greatly increased or decreased the odds ratio (OR) of any variable. As disease complexes change with age and vary among types of production systems, the data was analyzed for each of these production variables, and separate models were constructed.17 Initial screening and model fitting were again performed for each age and type of production system. To obtain ORs and P values for the serological and lung-score categories coded as “low,” data for these variables were coded in reverse (0’s re-coded to 2’s, 1’s remained the same, and 2’s re-coded to 0’s). Models were evaluated for fit by plotting the residuals using Minitab software (State College, Pennsylvania). Variables with P values < .05 were retained in the models.
Results
Overall model
When the data was analyzed for the overall association of PRRSV-positive pigs and various pathogens, lesions, and serological profiles, SIV was 2.40 times as likely to be identified by PCR in PRRSV-positive pigs than in PRRSV-negative pigs (95% CI, 1.07-5.39). Also, PRRSV-positive pigs were more likely than PRRSV-negative pigs to have lung scores in the mid-range (OR, 1.65; 95% CI, 1.06-2.57), PRRS ELISA S:P ratios in the mid-range (OR, 9.37; 95% CI, 4.58-19.18), and PRRS ELISA S:P ratios in the high-range (OR, 4.65; 95% CI, 2.46-8.79). Age was a major confounder and was thus retained in the overall model.
Early-nursery model
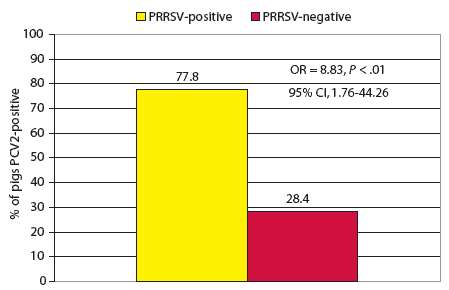
In the 3-week-old disease-interaction model, PCV2 was 8.83 times as likely to be identified by PCR in the PRRSV-positive pigs as in the PRRSV-negative pigs (Figure 1). Management system and disease severity were not confounders in this model.
| Figure 1: Co-infection model for porcine reproductive
and respiratory syndrome virus (PRRSV) and porcine circovirus type 2 (PCV2)
in 3-week-old pigs (n = 157) either positive for PRRSV by polymerase chain reaction (PCR) and with lung lesions consistent with PRRSV infection (9 pigs) or PRRSV-negative by these criteria (148 pigs). Pigs were tested for PCV2 by PCR on tissue samples at necropsy.
|
Late-nursery model
At 9 weeks of age, PRRSV-positive pigs, compared to PRRSV-negative pigs, were 15.57 times more likely than to be PCR-positive for SIV (95% CI, 1.77-137.23), 15.54 times more likely to have mid-range PRRSV S:P ratios (95% CI, 4.80-50.31), and 6.04 times more likely to have high-range PRRSV S:P ratios (95% CI, 2.57-14.19).
Early-mid-finishing and late-finishing models
At 16 weeks of age, PRRSV-positive pigs, were more likely than PRRSV-negative pigs, to be PCR-positive for SIV and to have mid-range PRRSV S:P ratios (Table 1). At 24 weeks of age, PRRSV-positive pigs were significantly less likely to be co-infected with Salmonella serovars (OR, 0.15; 95% CI, 0.04-0.61). However, PRRSV-positive pigs 24 weeks of age were more likely to have M hyopneumoniae S:P ratios in the mid-range (OR, 3.36; 95% CI, 1.11-10.24 ) or high range (OR, 2.70; 95% CI, 1.03-7.12) and SIV H1N1 S:P ratios in the mid-range (OR, 4.78; 95% CI, 1.84-12.38).
Table 1: Disease interaction model for porcine reproductive and respiratory syndrome virus (PRRSV) and swine influenza virus (SIV) in 9-week-old (n = 149) and 16-week- old pigs (n = 152) that were either positive or negative for PRRSV*
* PRRSV-positive pigs were positive by polymerase chain reaction (PCR) on tissue samples and had lung lesions consistent with PRRSV infection; PRRSV-negative pigs were negative by these criteria. † P values obtained from the logistic regression model. ‡ Tissue samples were tested by PCR for swine influenza virus (SIV) type A. § Serum samples were tested by ELISA for antibodies to PRRS virus, with sample:positive (S:P) results categorized as ≤ 0.49 (low), 0.50 to 0.99 (medium), and ≥ 1.00 (high). |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||
One-site model
When the data were stratified by type of management system, PRRSV-positive pigs in the one-site management system model were 0.21 times as likely as PRRSV-negative pigs to be culture-positive for H parasuis (95% CI, 0.06-0.78). Also, PRRSV-positive pigs had significantly greater associations with lung scores in the mid-range (OR, 2.70; 95% CI, 1.04-7.00) and high range (OR, 4.98; 95% CI, 1.19-20.76), were more likely to have PRRSV S:P ratios in the mid-range (OR, 16.37; 95% CI, 4.21-63.57) and high range (OR, 4.55; 95% CI, 1.51-13.66), and were 0.35 times as likely to have SIV H1N1 S:P ratios in the mid-range (95% CI, 0.13-0.95). Disease severity and age were confounding factors and were thus retained in this model.
Two-site model
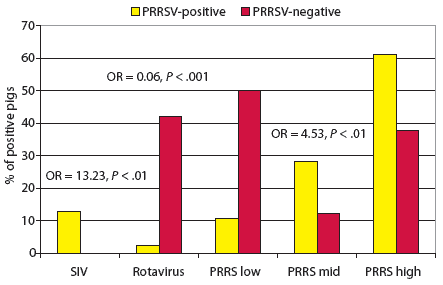
In the two-site management system model (Figure 2), rotavirus infection was less likely to be found in PRRSV-positive pigs than in PRRSV-negative pigs. However, PRRSV S:P ratios in the mid-range were significantly associated with PRRSV-positive pigs. In the bivariate analysis, there was a significant association of the outcome variable “PRRSV-positive” with the exposure variable “PCR-positive for SIV type A” (OR, 13.23); however, it could not be included in the disease model because none of the PRRSV-negative pigs were positive for SIV, and a value of 0 cannot be utilized in this type of model construction.
| Figure 2: Co-infection model for porcine reproductive
and respiratory syndrome virus (PRRSV) for two-site management systems
(n = 175). Pigs 3 to 24 weeks old were considered PRRSV-positive if tissue
samples at necropsy were positive by polymerase chain reaction (PCR) and
if lung lesions were consistent with PRRSV infection. Pigs not meeting
these criteria were considered PRRSV-negative. Sera were tested by PRRS
ELISA, with results reported as sample:positive (S:P) ratios in three ranges:
low (< 0.49), mid (0.50-0.99), or high ( > 1.0). Tissue
samples were tested by PCR at necropsy for swine influenza virus type A
(SIV), and fecal samples were examined by electron microscopy for porcine
rotavirus. The SIV variable was not included in the model because of a
0 value in the equation. The odds ratio (OR) and P value for SIV
were obtained using bivariate analysis. All other ORs and P values
were obtained from the logistic regression model.
|
Three-site model
In the three-site management system model, PRRSV-positive pigs were 10.09 times more likely than PRRSV-negative pigs to have PRRS ELISA S:P ratios in the mid-range (95% CI, 3.16-32.24), 5.04 times more likely than PRRSV-negative pigs to have PRRS ELISA S:P ratios in the high-range (95% CI, 1.94-13.06), and 3.37 times more likely than PRRSV-negative pigs to have SIV H1N1 S:P ratios in the mid- range (95% CI, 1.54-7.43).
Discussion
This epidemiologic analysis in an integrated production system confirms that there are age-related and management system-related pathogen associations of PRRSV co-infection. The most common co-infector in PRRSV-positive pigs was SIV, with SIV exposure variables significantly associated with the outcome variable “PRRSV-positive” in seven of the eight models constructed. Other common associations among the models included higher lung scores and higher S:P ratios in ELISAs for PRRS, M hyopneumoniae, and SIV H1N1. In the overall model, as expected, age was a major confounder. However, type of management system was not, which was surprising, considering that different types of management systems are known to change the dynamics of many diseases.18
When the data were analyzed in the 3-week-old model, PCV2 was the only remaining variable significantly associated with the outcome variable “PRRSV-positive” after construction and fitting of the final model. Although PCV23 and porcine circovirus associated disease (PCVAD) manifest themselves in the late-nursery to mid-finishing stages,3,17 these data suggest that efforts for prevention and control need to be focused on the sows from gestation through lactation. Among the 157 pigs included in this model, only nine were PCR-positive for PRRSV and had lesions suggestive of PRRSV infection; however, seven of these nine pigs (78%) were co-infected with PCV2, compared with approximately 28% of the PRRSV-negative pigs. As previously suggested,4 young pigs co-infected with PCV2 and PRRSV, and either clinically or subclinically affected, may act as a source of infection for other animals, and may become a serious risk to their penmates in the nursery as their maternal antibodies decline. This may in turn increase the prevalence of PRDC or PCVAD in the late-nursery to mid-finishing stages where these syndromes tend to manifest themselves most commonly.
The 9-week-old and 16-week-old pigs had very similar profiles in their respective models, but with varying degrees of association. In both the 9-week-old and 16-week-old groups, PRRSV-positive pigs were more likely than PRRSV-negative pigs to be co-infected with SIV and to have higher PRRSV S:P ratios, with the associations slightly stronger in the 9-week-old group. These results suggest that SIV plays a role as a statistically significant co-infector with PRRSV in the late nursery and early- to mid-finishing stages.
The 24-week-old PRRSV-positive pigs were significantly less likely than the PRRSV-negative pigs to be co-infected with Salmonella serovars. This might have been due to an up-regulation or modification of the immune system towards effector cells that destroy invasive bacteria.3,4,14 In addition, the PRRSV-positive pigs in this group had significantly higher ELISA S:P ratios for M hyopneumoniae and SIV H1N1. Higher titers to SIV would be expected in this group, as there was a significant association with SIV co-infection in PRRSV-positive pigs in the 9-week-old and 16-week-old models. However, 24-week-old PRRSV-positive pigs were not more likely than PRRSV-negative pigs to be PCR-positive for M hyopneumoniae. This finding may indicate that M hyopneumoniae was indeed a statistically significant co-factor in PRRSV-positive pigs prior to 24 weeks of age, but was missed due to the cross-sectional nature of this study.
In the one-site management system model, PRRSV-positive pigs were more likely to have lung scores in the 11% to 50% and 51% to 100% ranges, in addition to having very strong associations with mid- and high-range PRRSV S:P ratios. The associations between higher S:P ratios and higher lung scores were greater here than in any other model, possibly due to the constant circulation of virus that occurs in farrow-to-finish production facilities. Systems that do not experience an acute outbreak of PRRSV may facilitate a constant risk of exposure to naive pigs through fomite, aerosol, or nose-to-nose contact. Fomites likely play a large role in farrow-to-finish facilities because, even though there is supposed to be a division of labor between the various age groups, this may not be practiced faithfully. Haemophilus parasuis, on the other hand, was less likely to be found in PRRSV-positive pigs than in PRRSV-negative pigs in one-site facilities. This is consistent with previous reports that suggest that PRRSV does not potentiate the clinical course of H parasuis, or that PRRSV may even stimulate the phyagocytotic activity of alveolar macrophages, thereby decreasing the bacterial load in the lung.14,15
In the two-site management systems, PRRSV S:P ratios in the 0.50 to 0.99 range were more likely to be found in PRRSV-positive pigs than in PRRSV-negative pigs, as in the other age-group and management-system models. Although SIV was not included in the model, diagnostically speaking, it should be considered an important co-infector in the two-site management systems. The perplexing result in this model is the very strong association of rotavirus with PRRSV-negative pigs. This may be a result of an up-regulation of macrophages in the intestinal mucosa when these pigs are young, which may have a protective effect.
In the three-site management systems, there was no association between PRRSV-positive status and statistically significant co-infections with agents identified either by PCR or culture. However, there was a significant association between PRRSV-positive status and SIV H1N1 S:P ratios in the 0.50 to 0.99 range, which may suggest a variation in response or clinical outcome of exposure to SIV in PRRSV-positive pigs. Mid- and high-range PRRSV S:P ratios were strongly associated with PRRSV-positive pigs, as in all models.
Overall, the dynamics of PRRSV infection, co-infection, and disease manifestation changed with age and type of management system, as hypothesized. However, in most models, PRRSV-positive status was associated with more lung damage, SIV co-infection, and higher SIV H1N1, M hyopneumoniae, and PRRS ELISA S:P ratios. Research is needed to further investigate the apparently “protective” effect that PRRSV seems to have against H parasuis, Salmonella serovars, and rotavirus. The strong association between the variables “PRRSV-positive” and “PCV2-positive by PCR” in pigs 3 weeks of age is a major concern regarding the immunocompetence of nursery pigs. The nursery pig’s immune profile and ability to respond to vaccines (eg, for SIV, M hyopneumoniae, Salmonella) in the presence of PRRSV and PCV co-infection should be evaluated.
Implications
- Manifestation of PRRS and associated co-factors change with age and with different types of management systems.
- Under the conditions of this study in one-, two- and three-site management systems, PCV2 is a statistically significant co-infector in PRRSV-positive pigs 3 weeks of age.
- Under the conditions of this study, SIV is a consistent co-infector in PRRSV-positive pigs.
- Further research is warranted on disease caused by co-infection of pathogens and efficacy of vaccines in preventing disease caused by multiple pathogens.
- Timing of vaccination for disease complexes associated with PRRSV needs to be further investigated, especially when multiple antigens must be presented at once.
References
1. Albina E. Epidemiology of porcine reproductive and respiratory syndrome (PRRS): an overview. Vet Microbiol. 1997;55:309–316.
2. Neumann EJ, Kliebenstein JB, Johnson CD, Mabry JW, Bush EJ, Seitzinger AH, Green AL, Zimmerman JJ. Assessment of the economic impact of porcine reproductive and respiratory syndrome on swine production in the United States. JAVMA. 2005;227:385–392.
*3. Thacker EL. Immunology of the porcine respiratory disease complex. Vet Clin North Am Food Anim Pract. 2001;17:551–565.
4. Dorr PM, Baker RB, Almond GW, Wayne SR, Gebreyes WG. Epidemiologic assessment of porcine circovirus type 2 coinfection with other pathogens in swine. JAVMA. 2007;230:244–250.
5. Brockmeier SL, Palmer MV, Bolin SR, Rimler RB. Effects of intranasal inoculation with Bordetella bronchiseptica, porcine reproductive and respiratory syndrome virus, or a combination of both organisms on subsequent infection with Pasteurella multocida in pigs. Am J Vet Res. 2001;62:521–525.
6. Wills RW, Gray JT, Fedorka-Cray PJ, Yoon KJ, Ladely S, Zimmerman JJ. Synergism between porcine reproductive and respiratory syndrome virus (PRRSV) and Salmonella choleraesuis in swine. Vet Microbiol. 2000;71:177–192.
7. Galina L, Pijoan C, Sitjar M, Christianson WT, Rossow K, Collins JE. Interaction between Streptococcus suis serotype 2 and porcine reproductive and respiratory syndrome virus in specific pathogen-free piglets. Vet Rec. 1994;134:60–64.
8. Feng W, Laster SM, Tompkins M, Brown T, Xu JS, Altier C, Gomez W, Benfield D, McCaw MB. In utero infection by porcine reproductive and respiratory syndrome virus is sufficient to increase susceptibility of piglets to challenge by Streptococcus suis type II. J Virol. 2001;75:4889–4895.
9. Harms PA, Sorden SD, Halbur PG, Bolin SR, Lager KM, Morozov I, Paul PS. Experimental reproduction of severe disease in CD/CD pigs concurrently infected with type 2 porcine circovirus and porcine reproductive and respiratory syndrome virus. Vet Pathol. 2001;38:528–539.
10. Roberts NE, Almond GW. Infection of growing swine with porcine reproductive and respiratory syndrome virus and Mycoplasma hyopneumoniae—effects on growth, serum metabolites, and insulin-like growth factor-I. Can Vet J. 2003;44:31–37.
11. Van Reeth K, Nauwynck H, Pensaert M. Dual infections of feeder pigs with porcine reproductive and respiratory syndrome virus followed by porcine respiratory coronavirus or swine influenza virus: a clinical and virological study. Vet Microbiol. 1996;48:325–335.
12. Van Reeth K, Nauwynck H, Pensaert M. Clinical effects of experimental dual infections with porcine reproductive and respiratory syndrome virus followed by swine influenza virus in conventional and colostrum-deprived pigs. J Vet Med B Infect Dis Vet Public Health. 2001;48:283–292.
13. Thacker EL, Halbur PG, Ross RF, Thanawongnuwech R, Thacker BJ. Mycoplasma hyopneumoniae potentiation of porcine reproductive and respiratory syndrome virus-induced pneumonia. J Clin Microbiol. 1999;37:620–627.
14. Segales J, Domingo M, Balasch M, Solano GI, Pijoan C. Ultrastructural study of porcine alveolar macrophages infected in vitro with porcine reproductive and respiratory syndrome (PRRS) virus, with and without Haemophilus parasuis. J Comp Pathol. 1998;118:231–243.
15. Cooper VL, Doster AR, Hesse RA, Harris NB. Porcine reproductive and respiratory syndrome: NEB-1 PRRSV infection did not potentiate bacterial pathogens. J Vet Diagn Invest. 1995;7:313–320.
16. Van Alstine WG, Stevenson GW, Kanitz CL. Porcine reproductive and respiratory syndrome virus does not exacerbate Mycoplasma hyopneumoniae infection in young pigs. Vet Microbiol. 1996;49:297–303.
17. Segales J, Allan GM, Domingo M. Porcine circovirus diseases. Anim Health Res Rev. 2005;6:119–142.
18. Harris DL. Multi-Site Pig Production. Ames, Iowa: Iowa State University Press; 2000.
* Non-refereed reference.