Case study |
Peer reviewed |
A cluster of farms experiencing severe porcine circovirus associated disease: Clinical features and association with the PCV2b genotype
Un grupo de granjas que experimentaron enfermedad severa asociada a circovirus porcino: Características clínicas y asociación con el genotipo PCV2b
Maladie sévère associée au circovirus dans un regroupement de fermes: Caractéristiques cliniques et association avec le génotype PCV2b
Kyle P. Horlen, DVM; Paula Schneider; Joseph Anderson; Jerome C. Nietfeld, DVM, MS, PhD, Diplomate ACVP; Steven C. Henry, DVM, Diplomate ABVP; Lisa M. Tokach, DVM, Diplomate ABVP; Raymond R. R. Rowland, MA, PhD
KPH, PS, JA, JCN, RRRR: Department of Diagnostic Medicine and Pathobiology, College of Veterinary Medicine, Kansas State University, Manhattan, Kansas. SCH, LMT: Abilene Animal Hospital, PA, Abilene, Kansas. Corresponding author: Dr Kyle P. Horlen, Department of Diagnostic Medicine/Pathobiology, 1800 Denison Ave K-212, Manhattan, KS 66506–5601; Tel: 785–532–4855; Fax: 785–532–4039; E-mail: horlen.dvm@gmail.com
Cite as: Horlen KP, Schneider P, Anderson J, et al. A cluster of farms experiencing severe porcine circovirus associated disease: Clinical features and association with the PCV2b genotype. J Swine Health Prod. 2007;15(5):270–278.
Also available as a PDF.
SummaryThe objective of this study was to characterize the clinical, histopathological, and virological features of an outbreak of severe porcine circovirus associated disease (PCVAD) on four farms located in the same geographical region of Kansas. The outbreak of PCVAD was evident by increased mortality and the appearance of postweaning multisystemic wasting syndrome, porcine dermatitis and nephropathy syndrome, and neurological signs. Whole porcine circovirus type 2 (PCV2) genomes were derived by sequencing polymerase chain reaction (PCR) products amplified from tissues. Nucleotide differences between North American and European isolates were used to construct a genotype-specific PCR assay for detection of different genotypes. Genetic sequencing of PCV2 from 12 PCVAD-affected pigs showed that all viruses were associated with the PCV2b genotype and were distant from historical PCV2a isolates. Routine virus isolation from tissues of PCVAD-affected pigs yielded both PCV2a and PCV2b viruses, as well as a porcine teschovirus.The identification of PCV2b in Kansas is consistent with the increased incidence of both severe PCVAD and variant PCV2 genotypes worldwide. The ultimate source of the PCV2b virus within the production systems studied remains unclear. Genotype-specific PCR assays provide new tools for understanding the contributions of PCV2a and PCV2b viruses to PCVAD. | ResumenEl objetivo de este estudio fue determinar las características clínicas, histopatológicas, y virológicas de un brote de enfermedad severa asociada a circovirus porcino (PCVAD por sus siglas en ingles) en cuatro granjas localizadas en la misma región geográfica de Kansas. El brote de PCVAD fue evidente por el aumento de mortalidad y la aparición del síndrome de desgaste multisistémico post destete, dermatitis porcina y síndrome de neuropatía, y signos neurológicos. Los genomas completos del circovirus porcino tipo 2 (PCV2 por sus siglas en inglés) se derivaron de la secuenciación de la amplificación de los productos de la reacción en cadena de la polimerasa (PCR por sus siglas en inglés) de tejidos. Se utilizaron las diferencias entre los aislamientos Norteamericanos y Europeos para construir un PCR específico de genotipo para la detección de los diferentes genotipos. Las secuencias del PCV2 de los 12 cerdos afectados con el PCVAD demostraron que todos los virus estaban asociados al genotipo PCV2b y que eran distantes del aislamiento PCV2a. El aislamiento rutinario de tejidos de cerdos afectados con PDCAV mostraron los dos genotipos PCV2a y PCV2b asi como el virus tescho de puercos. La identificación de PCV2b in Kansas es consistente con el incremento de la incidencia severa del os dos PCVAD genotipos y la variante del genotipo PCV2 a nivel mundial.. La fuente del PCV2b en el sistema de producción permanece desconocida. El PCR específico de genotipo provee una herramienta novedosa para entender la contribución del PCV2a y PCV2b en la PCVAD. | ResuméL’objectif de la présente étude était de déterminer les caractéristiques cliniques, histopathologiques, et virologiques d’une poussée de cas sévères de maladie associée au circovirus porcin (PCVAD) sur quatre fermes localisées dans la même région géographique du Kansas. La poussée de cas était manifeste par une augmentation de la mortalité et l’apparence d’un syndrome de dépérissement multi-systémique post-sevrage, de dermatite porcine et d’un syndrome néphropatique, de même que de signes neurologiques. Le génome entier des circovirus de type 2 (PCV2) a été déduit en séquençant les produits des réactions d’amplification en chaîne (PCR) effectuées à partir des tissus. Des différences dans les séquences nucléotidiques entre les isolats nord-américains et européens ont été utilisées afin de construire une épreuve PCR spécifique de génotype afin de détecter les différents génotypes. Le séquençage génétique du PCV2 provenant de 12 porcs affectés par le PCVAD a démontré que tous les virus étaient associés avec le génotype PCV2b et étaient distants de l’isolat PCV2a historique. L’isolement viral de routine à partir des tissus de porcs affectés par PCVAD a permis d’obtenir les virus PCV2a et PCV2b, de même que qu’un teschovirus porcin. L’identification de PCV2b au Kansas est en accord avec l’augmentation d’incidence de PCVAD sévère et de variants génotypiques de PCV2 à travers le monde. La source ultime de PCV2b à l’intérieur des systèmes de production étudiés n’est pas clairement établie. Des essais de PCR spécifiques de génotype fournissent de nouveaux outils pour comprendre les contributions de PCV2a et PCV2b au PCVAD. |
Keywords: swine, porcine
circovirus associated disease, porcine circovirus type 2, PCV, polymerase
chain reaction, PCR, porcine teschovirus
Search the AASV web site
for pages with similar keywords.
Received: July
25, 2006
Accepted: March
22, 2007
Porcine circovirus type 2 (PCV2) is a small nonenveloped DNA virus with a single-stranded circular genome and is assigned to the family Circoviridae.1 Porcine circovirus type 2 is recognized as the primary etiological agent associated with a group of complex multi-factorial diseases classified as porcine circovirus associated disease (PCVAD), of which postweaning multisystemic wasting syndrome (PMWS) and porcine dermatitis and nephropathy syndrome (PDNS) are major syndromes.2 Postweaning multisystemic wasting syndrome was described in 1997 as a new disease seen in several high-health herds in western Canada3 and has become a growing concern in the swine industry throughout the world.4
Although it is now recognized in many pig-producing countries, the etiology of PDNS remains unclear. Porcine circovirus type 2 has been implicated as the causative agent, but the simple presence of PCV2 in most animals affected with PDNS is not conclusive proof of pathogenesis. However, the pathological changes of PDNS are consistent with an immune-complex disorder, and it has been hypothesized that extremely high PCV2 serum antibody titers and immune-complex deposition are a significant factor for disease.5 Phylogenetic analyses of PCV2 open reading frame 2 (ORF2) gene sequences show branches that are identified within different geographic regions.6,7 The virus strains of recent interest are those that are identified as PCV2a and PCV2b. The terminology “PCV2a” and “PCV2b” is based upon whether the isolated virus is more similar to historical North American isolates (“a”) or European isolates (“b”). On the basis of restriction endonuclease digestion, most PCV2 genotypes in the United States can be further described by the restriction fragment length polymorphism patterns 4–2–2 (PCV2–422 genotype) and 3–2–1 (PCV2–321 genotype), which, with some exceptions, correspond to strains PCV2a and PCV2b, respectively. Historically, PCVAD cases in the United States have been associated with viruses possessing the 422 genotype.8 Recent cases of PCVAD in Canada are linked to the emergence of the 321 genotype, which shows close similarities to certain PCV2b isolates, such as a 1998 French isolate (GenBank AF055393).8,9
The interactions between PCV2 and other factors, including infectious agents, are well documented. Both porcine parvovirus (PPV) and porcine reproductive and respiratory syndrome (PRRS) virus are considered cofactors in the development of PMWS.10,11 Additionally, bacteria such as Mycoplasma hyopneumoniae have been shown to increase the severity of clinical signs in a manner similar to PPV and PRRS virus.12 It is likely that numerous other infectious cofactors are involved in the etiology of PCVAD.
In this study, we identified four herds within the same geographical region of Kansas that were experiencing severe PCVAD. The term “severe” was used as a means to describe this outbreak as not only meeting the criteria of Sorden’s definition13 of PMWS, but also exhibiting a dramatic increase in the morbidity and mortality of growing pigs. Furthermore, severe PCVAD was observed in both serologically PRRS-negative and PRRS-positive herds and included some clinical signs not typically associated with PMWS or PDNS. The purpose of this study was to characterize the clinical, pathological, and virological features of an outbreak of severe PCVAD in Kansas, including the genotypic properties of PCV2 isolates recovered from affected and nonaffected pigs, possible sources of PCV2 in a production system, and the identity of other viral cofactors responsible for atypical clinical signs.
Case description
Herd history
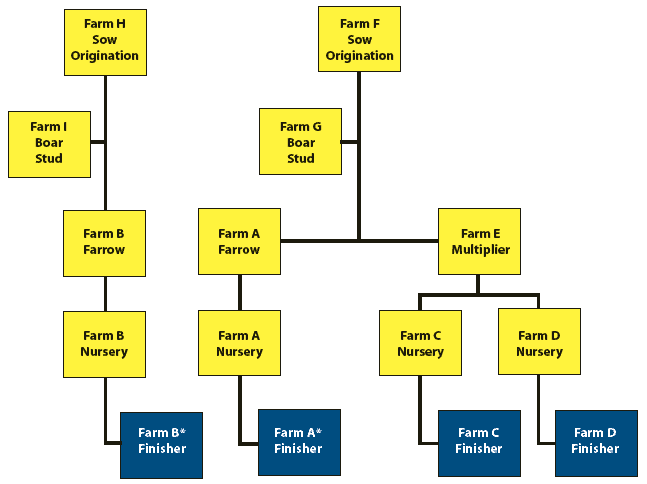
In Kansas, the first clinical recognition of severe PCVAD occurred in November 2005, affecting four separate finisher operations located in the northeastern part of the state. For the purpose of convenience, we labeled these sites Farms A, B, C, and D. As shown in Figure 1, Herds A and B were PRRS-positive multi-site farrow-to-finish operations. Herds C and D were PRRS-negative, and served as nursery and finisher sites for weaned pigs from Herd E, a PRRS-negative sow farm. The PRRS status of each farm had been determined by routine monitoring for the presence of PRRS virus antibody using a standard PRRS ELISA (Idexx, Westbrook, Maine). Herds A and E received replacement gilts as weaned piglets from Herd F, a PRRS-negative multiplier, and semen from Herd G. Thus, the genetic source of Herds A, C, and D was the same. Herd B received animals from a different boar stud and gilt multiplier. Additionally, a fifth herd in the same area was investigated as possibly experiencing an outbreak of severe PCVAD following reports of increased morbidity and mortality. However, extensive investigation of several animals revealed no evidence of PCVAD despite the isolation of PCV2 from two pigs. Finally, a small farrow-to-finish herd that received semen from Farm G was investigated as experiencing severe PCVAD. Following confirmation of clinical signs and histopathologic lesions, samples from this farm, as well as samples from Farms A through D, were used to evaluate a set of differential PCV2 polymerase chain reaction (PCR) primers. All farms were considered to be of overall good health with minimal incidence of common pathogens such as swine influenza virus, Lawsonia intracellularis, and Mycoplasma hypneumoniae prior to the study investigation. For the initial investigation, approximately 20 pigs were sampled per site.
| Figure 1: Source and distribution flow chart in
a study of severe porcine circovirus associated disease (PCVAD) in four
separate finishers in the same geographical area. The gray boxes indicate
farm sites where PCVAD-positive pigs were found. White boxes identify the
sources of the affected pigs. Asterisks identify herds seropositive for
porcine reproductive and respiratory syndrome.
|
Clinical and pathological features of pigs from the four Kansas farms
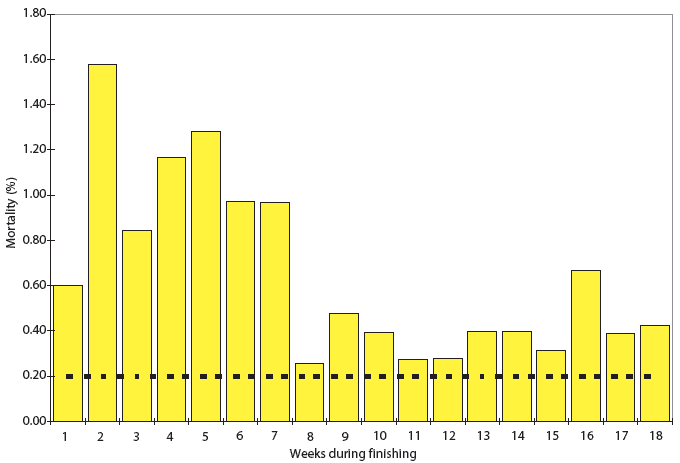
The first indication of a problem in herds A through D was the appearance of increased morbidity and mortality during finishing. Pigs entered the finisher at 9 to 10 weeks of age. Historically, mortality in these herds ranged from 3% to 5% during the finishing period, but had suddenly increased to as much as 15%. An example of the mortality profile for Farm A is presented in Figure 2. Detection of clinical signs and increased mortality led to an in-depth investigation at each farm. Most affected pigs could be placed into one of two groups. Pigs in one group had clinical signs and pathologic lesions compatible with PMWS, and pigs in the second group had clinical signs and pathologic lesions compatible with PDNS.
| Figure 2: Mortality in Herd A by week post placement.
The data include 6356 pigs in five PCVAD-affected lots. The dotted line
shows the historic mortality, which averaged 0.2% deaths per week.
|
Pigs in the PMWS group were noticeably smaller than their penmates and had long rough hair coats. At necropsy, the submandibular, inguinal, and bronchial lymph nodes were swollen. The lungs were wet and did not fully collapse, and, in most cases, there were areas of anterior-ventral pulmonary consolidation compatible with bacterial bronchopneumonia. A few pigs were icteric and their livers were slightly nodular and firmer than normal when incised. Microscopically, lymphocytes in germinal centers of the lymph nodes and tonsils and in the periarterial lymphatic sheaths of the spleen were consistently replaced by large macrophage-like cells and occasional multinucleated giant cells. In addition, in many pigs, lymphocytes in the paracortical areas of the lymph nodes and tonsil were also replaced by large macrophage-like or stromal cells. In areas of the lung without bronchopneumonia, there were interstitial infiltrates of macrophages and lymphocytes. This interstitial pneumonia was more severe in serologically PRRS-positive pigs. The kidneys typically contained multifocal, coalescing interstitial infiltrates of lymphocytes and plasma cells. In a few kidneys, multiple areas also contained large numbers of macrophages. Some tubules contained sloughed and fragmented cells. In the livers from the icteric pigs, there were multiple foci of necrosis, chronic inflammation, and portal fibrosis. Immunohistochemical (IHC) staining for PCV2 consistently revealed abundant antigen in the cytoplasm of macrophage-like cells in lymph nodes, tonsils, and spleen. In parenchymal organs, such as the lung, kidney, and liver, there was PCV2 antigen in macrophages and the nuclei of many epithelial cells, with positive staining present in almost all lungs and a minority of livers and kidneys.
Pigs in the PDNS group were in good body condition but had discrete red to purple to black papules involving the rear limbs, or large purple to black areas of epidermal discoloration over the entire body, or both. The inguinal lymph nodes were consistently markedly enlarged and hemorrhagic. The pleural and peritoneal cavities contained watery fluid (less than 1.0 L each) and the kidneys were swollen, wet, and usually had pinpoint hemorrhages covering the capsule. In several pigs, the spleen contained raised infarcts that measured approximately 1.0 to 2.5 cm in diameter. Microscopically, the kidneys’ glomeruli were swollen and filled with fibrin, and there was necrosis of the glomerular tufts. In most cases, all glomeruli were involved, but in a few pigs, less than half of the glomeruli were visibly damaged. Large numbers of tubules were filled with protein, hemoglobin, erythrocytes, and cellular detritus. In most kidneys, the glomerular lesions were acute, but in a few pigs the lesions were chronic, with the fibrin in glomeruli organized by fibroblast-like cells. Commonly, there also was extensive interstitial inflammation consisting primarily of lymphocytes and plasma cells and, in some cases, small infiltrates of neutrophils, with the inflammation being predominately in the medulla. In most kidneys, there was fibrinoid necrosis and perivascular cuffing of small and medium-sized arteries by lymphocytes and plasma cells. In sections of discolored skin there was acute coagulative necrosis of the epidermis with edema, hemorrhage, neutrophilic and eosinophilic inflammation, and vascular necrosis in the dermis. In the spleens of most pigs with PDNS, there were scattered arteries with fibrinoid necrosis. In a few instances, the necrotic arteries were surrounded by microscopic to macroscopic areas of coagulative necrosis. Follicles in lymph nodes and tonsils contained groups of large epitheloid macrophages and occasional multinucleated giant cells. In some lymph nodes, focal areas of coagulative necrosis, some of which contained mineral, appeared to involve follicles. The medullary and trabecular sinuses of many nodes were filled with hemorrhage and neutrophils. Fibrinoid necrosis of arteries was also present in rare sections of lymph nodes, the epicardium, and the capsule of the adrenal gland. Immunohistochemical staining revealed small amounts of PCV2 antigen in macrophages in follicles of the lymph nodes and tonsil of most pigs. Rare sections of kidney were IHC-positive for PCV2, and all sections of skin were IHC-negative. In a few cases, all tissues were IHC-negative for PCV2.
In three of the four affected herds, numerous pigs had rectal prolapse and weakness in the rear legs. Nonsuppurative inflammation was observed in the brains and spinal cords of these pigs, but IHC staining for PCV2 was negative.
Diagnostic investigation
Whole PCV2 genome sequencing and phylogenetic analysis
Whole-genome PCV2 sequences were obtained from 12 PCVAD-affected pigs from the four affected farms. One gram of lung and lymph node tissue was homogenized in 10 mL of minimum essential medium (MEM) and tissue debris was removed by centrifugation. Total DNA was isolated from the clarified medium using a QIAamp DNA blood mini kit (Qiagen, Valencia, California). Polymerase chain reaction and DNA sequencing of whole PCV2 genomes was performed according to published protocols.6 Two overlapping segments were amplified using primer pairs CV1F 1336–5’AGGGCTGTGGCCTTTGTTAC and CV2R 536–5’TCTTCCAATCACGCTTCTGC, respectively, and CV3F 453–5’TGGTGACCGTTGCAGAGCAG and CV4R 1525–5’ TGGGCGGTGGACATGATGAG, respectively. The reaction mixture included the DNA template, 1� PCR buffer, 1.25 mM MgCl2, 0.2 mM of each deoxyribonucleotide triphosphate, 0.2 uM each primer, and 2.5 U Taq polymerase. Polymerase chain reaction conditions were one cycle at 95�C for 4 minutes followed by 40 cycles at 93�C for 45 seconds, 50�C for 45 seconds, and 72�C for 90 seconds. At the end, the reaction was incubated at 72�C for 10 minutes. Deoxyribonucleic acid sequences were determined using the PCR products, the relevant primer sets, and a sequencing kit from Beckman-Coulter (Fullerton, California). Analyses were performed on a Beckman-Coulter CEQ 8000 sequencer. Prior to sequencing, the PCR products were purified using a Wizard SV Gel and PCR Clean-Up System (Promega, Madison, Wisconsin). Analysis of the sequence chromatograms and assembly of the sequences were performed using several computer-based programs, including 4-Peaks (Mekentosj, Amsterdam, Netherlands), Vector NTI (Invitrogen, Calsbad, California), Clustal-X (Statsbourg Bioinformatics, Illkirch-Graffenstaden, France), and Gene Jockey II (Biosoft, Cambridge, United Kingdom). Phylogenetic trees were constructed using the whole 1768 nucleotide genome for each isolate. Sequences were aligned using Clustal-X. The unrooted phylogram was generated by a neighbor-joining method using Mega (Biodesign Institute, Tempe, Arizona).14
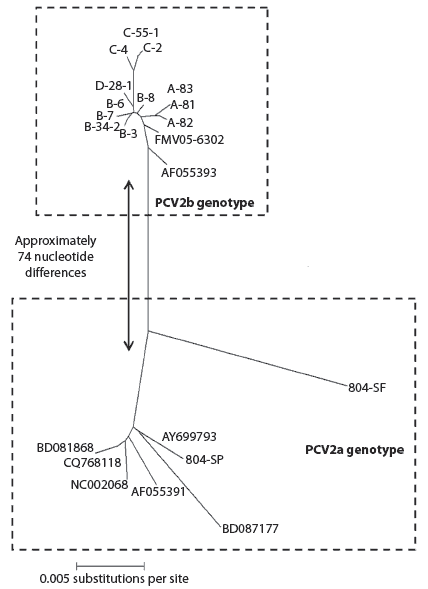
Within the phylogenetic tree, all 12 sequences formed a single group, which was located near the 1998 French PCV2b isolate, AF0553939 (Figure 3). This group of sequences was approximately 99.5% identical to AF055393. Sequences from a given farm tended to cluster together into the same sub-branch. A fifth herd located in the same general area was investigated as possibly experiencing an outbreak of PCVAD; however, even though the affected pigs were PCR-positive for PCV2, histopathology showed no evidence of PCVAD. Two PCV2 isolates were sequenced from two non-affected Kansas pigs. The sequences, identified as 804-SF and 804-SP in Figure 3, localized near other historical North American strains such as the PCV2a isolate AF055391. Sequences 804-SF and 804-SP were well separated from the PCV2 sequences of other Kansas PCVAD-affected pigs. The 12 Kansas PCV2 sequences isolated from PCVAD-affected pigs were only approximately 94% identical to AF055391. Differences at the nucleotide level translate into amino acid differences. Analysis of a representative PCV2b Kansas isolate and the Kansas PCV2a isolate (804-SF) revealed 12 amino acid differences, which were located between amino acids 57 to 91 and 151 to 210. Seven of the amino acid changes were nonconserved.
| Figure 3: Phylogenetic analysis of Kansas porcine
circovirus type 2 (PCV2) isolates in an outbreak of severe porcine circovirus
associated disease (PCVAD) on four Kansas finishers. An unrooted phylogram
shows locations of PCV2 isolates. The upper box shows isolates from Herds
A through D. Note the location of the French isolate, AF055393, and Canadian
isolate FMV05–6302. The lower box shows locations of two Kansas PCV2
isolates, 804-SP and 804-SF, from pigs showing no histological evidence
of PCVAD. Other isolates, identified by GenBank numbers, represent typical
American and Canadian isolates.
|
Selective PCR amplification of PCV2a and PCV2b genotypes in clinical samples
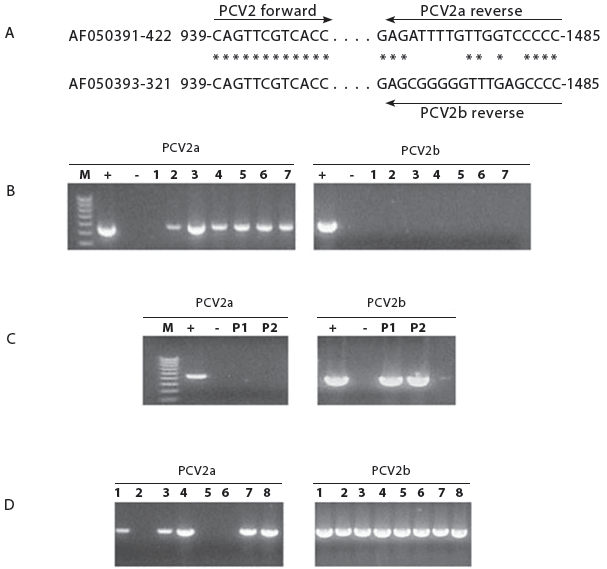
Utilizing genetic sequences unique to either PCV2a or PCV2b viruses, we designed genotype-specific primers that could be used to rapidly identify the different genotypes in tissues. As shown in Figure 4A, both primer sets contained the same forward primer, but different reverse primers. The original PCV2a-specific PCR-specific assay was performed according to Kim et al.15 Deoxyribonucleic acid was amplified using 35 cycles at 94�C for 30 seconds, 60�C for 30 seconds, and 72�C for 60 seconds, followed by a 10-minute extension at 72�C. The PCR products were electrophoresed on a 2% agarose gel and visualized under UV light after staining with ethidium bromide. Each run included positive and negative controls.
| Figure 4: Polymerase chain reaction (PCR) amplification
of two genotypes of porcine circovirus (PCV), PCV2a and PCV2b, in clinical
samples from a severe outbreak of porcine circovirus associated disease
(PCVAD) in four finishers in Kansas. A: Primers used to amplify PCV2a and
PCV2b genotypes; B: PCR results for seven boars from Herd G, with none
positive for PCV2b; C: PCR results for two pools (P1 and P2, three pigs/pool)
containing lymph node and lung tissue from pigs with severe PCVAD. Both
P1 and P2 were positive only for PCV2b; D: PCV2 screening of eight virus
isolates from pigs from Farms A through D with severe PCVAD, showing five
isolates positive for both PCV2a and PCV2b. In all frames,
“M” is a 1KB DNA marker, “+” is a positive control
of cloned template of PCV2a or PCV2b, and “-”
is a negative control.
|
Since PCV2 has been shown to be present in semen,16 the boar stud provides a logical source for the dissemination of new PCV2 strains. We sought to determine if the boar stud, identified as Farm G in Figure 1, could be the source of the PCV2b virus in Farms A, C, and D. Farm G contained approximately 300 boars with replacements sourced from two US farms. There was no evidence of PCVAD in the boar stud. Seven boars from Farm G, culled because of poor semen production, were removed and necropsied. Gross and microscopic analysis of lungs, lymph nodes, and kidneys confirmed that all boars were negative for PMWS and PDNS, and therefore, considered negative for PCVAD. Polymerase chain reaction amplification of lung and lymph node homogenates was performed using the PCV2a-specific and PCV2b-specific primers. The PCR results, presented in Figure 4B, showed that six of the seven boars were positive for PCV2a. However, all boars tested were negative for PCV2b. The PCR products were sequenced to confirm the presence of only PCV2a DNA. In contrast, PCR of pooled tissues from six PCVAD-affected pigs from a farm that received semen from Farm G showed no evidence of PCV2a sequence, but were positive for PCV2b (Figure 4C). These results, along with Figure 3, support the association of PCV2b with PCVAD-affected pigs, but fail to identify the boar stud as the PCV2b source.
We also used the PCV2a and PCV2b primers to amplify DNA from eight PCV2 isolates obtained by virus isolation from tissues of pigs that had originated from Farms A through D and that were severely affected with PCVAD. Isolation of PCV2 was performed on monolayers of rapidly dividing swine testicular (ST) cells (approximately 50% confluent). Cells were maintained in MEM supplemented with 7% fetal bovine serum and antibiotics (penicillin, streptomycin, amphotericin B, and ciprofloxacin). Monolayers were incubated with different dilutions of clarified homogenates prepared from lung, lymph node, and kidney collected from affected pigs. After 3 days, the media containing the virus was removed and stored at -80�C. To detect the presence of virus, monolayers were fixed in 80% aqueous acetone and stained with FITC-labeled porcine anti-PCV2 antibody (VMRD, Pullman, Washington). Cells were incubated with antibody for 1 hour, extensively washed with phosphate buffered saline (PBS), counterstained with Evans blue, and then viewed under a fluorescence microscope. Polymerase chain reaction analysis revealed that all isolates were positive for PCV2b. Interestingly, five of the isolates were positive for both PCV2a and PCV2b (Figure 4D). Whole-genome sequences, obtained using the method described for Figure 3, showed only the presence of PCV2b sequence.
Detection of porcine teschovirus (PTV) and porcine enterovirus (PEV)
Several of the PCVAD-affected pigs had neurological lesions that were not associated with the presence of PCV2 antigen. One possibility was that lesions were the result of a PTV or PEV infection. Polymerase chain reaction to detect PTV and PEV was performed according to the technique described by Palmquist et al.17 Total RNA was extracted from tissue homogenates composed of lung, lymph node, and spinal cord using a Qiagen RNAeasy kit (Valencia, California). Reverse transcriptase PCR was performed using a Qiagen One Step RT-PCR kit, according to the manufacturer’s recommendations. Forward and reverse primers were PTV-F 5’GTGGCGACAGGGTACAGAAGAG and PTV-R 5’GGCCAGCCGCGACCCTGTCAG, respectively. The PCR reaction conditions were 40 cycles at 94�C for 60 seconds, 56�C for 60 seconds, and 72�C for 60 seconds, followed by a 10-minute extension at 72�C. The PCR primers are designed to yield a 163-bp product for PTV and a 180-bp product for PEV.
Amplification of RNA from tissues of PCVAD-affected animals identified a 163-bp product in almost all pigs. Rarely, we found both PTV and PEV PCR products, but never the 180-bp PEV product alone. The PTV product was also obtained from tissues of pigs that showed no clinical or histological evidence of PCVAD. A PTV was isolated on ST cells from lung, lymph node, and spinal cord tissue homogenate of a PCVAD-affected pig on Farm C. To further subtype the PTV, we completed a second PCR reaction incorporating the degenerate forward and reverse primers, 2163F 5-GGCAATGCAAGGYCATTATGT and 2746R 5’AGCCCRCAYCTCADRTARGTGGCATG, respectively, under amplification conditions as described above. The PCR product was sequenced and then compared with known PTV sequences from GenBank. Analysis of this PTV sequence revealed that it is most closely related to a group of PTV-6 isolates.
Discussion
Severe PCVAD in a group of Kansas herds is a disease problem not previously encountered in this region. The isolation of PCV2 isolates from PCVAD-affected pigs that were clearly distinct from historical US isolates indicates that PCVAD-associated PCV2 isolates on Farms A through D clearly belonged to the PCV2b genotype. Since we have no historical information on PCV2 sequences in the four Kansas herds prior to the first appearance of PCVAD, we cannot determine the temporal association between the introduction of PCV2b and the current PCVAD outbreak. Interestingly, the data that has emerged from the study of the Kansas herds paints a picture similar to data reported from Canada that links PCV2–321 with the increased incidence and severity of PCVAD. For example, the Animal Health Laboratory at the University of Guelph in Ontario, Canada, reported only one case of PCV2–321-associated PCVAD in 2004, whereas 135 cases were reported in 2005.18 Furthermore, the increased incidence of PCV2–321 is associated with an overall increase of PCVAD in Canada.19 When Kansas PCV2b isolates are compared to Canadian isolate FMV05–6302,20 greater than 99% similarity is noted.
Reasons for the association of PCV2b with increased severity of clinical disease are unclear. One possibility is that the PCV2b genotype possesses greater virulence, or perhaps, the PCV2b virus can escape existing herd immunity stimulated by previously circulating PCV2a strains. The 233 amino-acid capsid protein of PCV2 is translated from ORF2. Bioinformatic and structural analysis of the PCV2 genome and proteins indicate that the capsid protein is the principal component of the outer viral capsid, which interacts with the cell receptor on host cells and stimulates protective immunity.21,22 Current PCV2 vaccines are based on stimulating anti-capsid immunity. Analysis of the amino acid differences between representative Kansas PCV2a and PCV2b isolates revealed several potentially significant amino-acid differences. These data indicate that the nucleotide differences between the Kansas PCV2a and PCV2b viruses may translate into significant immunological and functional differences between the PCV2a and PCV2b capsid proteins. Experiments are currently underway to test the disease potential of the Kansas PCV2b isolates. Although we failed to identify a PCVAD-affected pig infected with PCV2a alone, it is likely that virulent strains do exist.
Although the focus of this paper is the association of PCV2b with severe PCVAD, PCV2b may also be present in pigs not affected by disease. Since three of the four affected farms received semen from the same source, we originally believed that the boar stud was the most logical source of the PCV2b genotype. However, analysis of a small number of healthy boars showed only the PCV2a genotype. Collectively, these data suggest that healthy boars can be a source of PCV2a. Additionally, even though PCV2b sequence was not found in the analysis of tissues from seven boars, we cannot rule out the possibility that boar studs are also harboring the PCV2b virus. However, the ability of the boars to shed PCV2 in semen was not studied. Both PCV2a and PCV2b genotypes were found in isolates from PCVAD-affected pigs, suggesting that pigs can be infected simultaneously with both genotypes. However, when isolates from dually infected pigs were sequenced, only PCV2b was detected. This observation suggests that the PCV2b genotype is the predominant population in affected pigs. The PCV2a virus is still present, but as a much smaller subpopulation. The presence of the two viruses within a single animal raises the intriguing possibility that the two PCV2 genotypes may undergo recombination. Studies addressing co-infection are in progress.
Porcine circovirus type-2 antigen has been found associated with brain and spinal cord lesions in PCVAD-affected pigs exhibiting neurological signs.23 However, in this study, we failed to identify PCV2 in association with the central nervous system lesions from the Kansas PCVAD-affected pigs. A search for teschoviruses in one PCVAD-affected pig yielded an isolate that was closely related to PTV-6. Although PTV-6 is a serogroup 1 enterovirus, it is not known to be associated with porcine enteroviral encephalomyelitis or Talfan disease as is PTV serogroup 1, serotype 1 (formerly PEV-1).24,25 The rare detection of PEV in clinical samples by PCR was judged not to be important in this instance. Additionally, isolations carried out in porcine kidney-15 cells resulted in no other viruses that were consistently detected among PCVAD-affected pigs. Overall, the contribution of teschoviruses and enteroviruses to PCVAD remains unclear. Multiple viruses have been implicated in PCVAD, and PTV may be only one of several contributing co-factors.
Since this study began, an additional six herds in the same geographical area have been diagnosed with severe PCVAD. Genotype-specific PCR has confirmed the presence of PCV2b in all affected herds (unpublished data). While many of the newly affected herds share genetic sources with Farms A, C, and D, not all herds with these genetic sources exhibit PCVAD. Therefore, the presence of PCVAD may be the result of introduction of PCV2b into the nurseries or finishing houses of the affected herds. Another possibility is that the affected farms may possess a unique combination of cofactors that works in concert with PCV2b infection. Elevated mortality and clinical signs of PCVAD have continued on affected farms and detection of PCV2b has remained constant. Efforts to control mortality have focused on early culling of affected pigs and vaccination, as well as implementation of the Madec 20-Point Plan that includes aspects of biosecurity, environmental management, and overall health, especially regarding possible co-infecting agents.26
Implications
- High mortality due to PCVAD in four Kansas herds infected with a PCV2b strain suggests a greater pathogenic potential in PCV2b isolates than in PCV2a isolates.
- Nucleotide differences between PCV2a and PCV2b present an opportunity to develop differential assays to further track PCV2 genotypes.
- As current PCV2 vaccines are based on stimulating capsid immunity, significant peptide differences between PCV2a and PCV2b capsid proteins might have implications on the immunological cross-protective capability of single-strain commercial vaccines.
- Even though PTV was found both in PCVAD-positive and PCVAD-negative pigs, it may represent a cofactor in disease development, particularly with respect to atypical clinical signs such as neurological disease.
References
1. Allan GM, Ellis JA. Porcine circovirus: a review. J Vet Diagn Invest. 2000;12:3–14.
2. Segales J, Allan GM, Domingo M. Porcine circovirus associated diseases. Anim Health Res Rev. 2005;6:119–142.
*3. Clark EG. Post-weaning multisystemic wasting syndrome. Proc AASP. Quebec, Canada. 1997;499–501.
4. Chadd SA. The characteristics and attempted control of post-weaning multisystemic wasting syndrome (PMWS) and porcine dermatitis and nephropathy syndromes (PDNS) in European pig herds – a review. Pig J. 2003;51:13–25.
5. Wellenber GJ, Stockhofe-Zurwieden N, de Jong MF, Boersma WJA, Elbers ARW. Excessive porcine circovirus type 2 antibody titers may trigger the development of porcine dermatitis and nephropathy syndrome: a case-control study. Vet Microbiol. 2004;99:203–214.
6. Fenaux M, Halbur PG, Gill M, Toth TE, Meng X. Genetic characterization of type 2 porcine circovirus (PCV-2) from pigs with postweaning multisystemic wasting syndrome in different geographic regions of North America and development of a differential PCR-restriction fragment length polymorphism assay to detect and differentiate between infections with PCV-1 and PCV-2. J Clin Microbiol. 2000;38:2494–2503.
7. de Boisseson C, Beven V, Bigarre L, Thiery R, Rose N, Eveno E, Madec F, Jestin A. Molecular characterization of porcine circovirus type 2 isolates from post-weaning multisystemic wasting syndrome-affected and non-affected pigs. J Gen Virol. 2004;85:293–304.
*8. Ellis JA, Harding J, Gangon C, Hamel A. Porcine circovirus associated disease in Canada: A circle within a wheel. Proc IPVS. Copenhagen, Denmark. 2006;23–34.
9. Meehan BM, McNeilly F, Todd D, Kennedy S, Jewhurst VA, Ellis JA, Hassard LE, Clark EG, Haines DM, Allan GM. Characterization of novel circovirus DNAs associated with wasting syndromes in pigs. J Gen Virol. 1998;79:2171–2179.
10. Krakowka S, Ellis JA, Meehan B, Kennedy S, McNeilly F, Allan G. Viral wasting syndrome of swine: experimental reproduction of postweaning multisystemic wasting syndrome in gnotobiotic swine by coinfection with porcine circovirus 2 and porcine parvovirus. Vet Pathol. 2000;37:254–263.
11. Harms PA, Sorden SD, Halbur PG, Bolin SR, Lager KM, Morozov I, Paul PS. Experimental reproduction of severe disease in CD/CD pigs concurrently infected with type 2 porcine circovirus and porcine reproductive and respiratory syndrome virus. Vet Pathol. 2001;38:528–539.
12. Opriessnig T, Thacker EL, Yu S, Fenaux M, Meng XJ, Halbur PG. Experimantal reproduction of postweaning multisystemic wasting syndrome in pigs by dual infection with Mycoplasma hyopneumoniae and porcine circovirus type 2. Vet Pathol. 2004;41:624–640.
13. Sorden SD. Update on porcine circovirus and postweaning multisystemic wasting syndrome (PMWS). Swine Health Prod. 2000;8:133–136.
14. Kumar S, Tamura K, Nei M. MEGA: molecular evolutionary genetics analysis software for microcomputers. Comput Appl Biosci. 1994;10:189–191.
15. Kim J, Choi C, Han DU, Chae C. Simultaneous detection of porcine circovirus type 2 and porcine parvovirus in pigs with PMWS by multiplex PCR. Vet Rec. 2001;149:304–305.
16. Larochelle R, Bielanski A, Muller P, Magar R. PCR detection and evidence of shedding of porcine circovirus type 2 in boar semen. J Clin Microbiol. 2000;12:4629–4632.
17. Palmquist JM, Munir S, Taku A, Kapur V, Goyal SM. Detection of porcine teschovirus and enterovirus type II by reverse transcription-polymerase chain reaction. J Vet Diagn Invest. 2002;14:467–480.
*18. Carmen S, McEwen B, DeLay J, Cai H, Fairles J. Porcine circovirus type 2-associated disease continued from fall of 2005. AHL Newsletter. 2006;10(1):6. Available at http://www.labservices.uoguelph.ca/units/ahl/documents/ANwsl10-1.pdf. Accessed 4 Apr 2007.
*19. Delay J, McEwen B, Carmen S, van Dreumel T, Fairles J. Porcine circovirus type 2-associated disease is increasing. AHL Newsletter. 2005;9(3):22. Available at http://www.labservices.uoguelph.ca/units/ahl/files/ANwsl9-3.pdf. Accessed 4 Apr 2007.
*20. Tremblay D, Fontaine G, Harel J, Elahi MS, Gagnon CA. Genetic characterization of porcine circovirus type 2 from pigs showing signs of post-weaning multisystemic wasting syndrome. Unpublished. Available at http://www.ncbi.nlm.nih.gov/entrez/viewer.fcgi?db=nucleotide&val=78172609. Accessed 1 May 2007.
21. Nawagitul P, Morozov I, Bolin SR, Harms PA, Sorden SD, Paul PS. Open reading frame 2 of porcine circovirus type 2 encodes a major capsid protein. J Gen Virol. 2000;81:2281–2287.
22. Cheung AK. Transcriptional analysis of porcine circovirus. J Virol. 2003;305:168–180.
23. Fenaux M, Halbur PG, Haqshenas G, Royer R, Thomas P, Nawagitgul P, Gill M, Toth TE, Meng XJ. Cloned genomic DNA of type 2 porcine circovirus is infectious when injected directly into the liver and lymph nodes of pigs: characterization of clinical disease, virus distribution, and pathologic lesions. J Virol. 2002;76:541–551.
24. Yamada M, Kozakura R, Ikegami R, Nakamura K, Kaku Y, Yoshii M, Haritani M. Enterovirus encephalomyelitis in pigs in Japan caused by porcine teschovirus. Vet Rec. 2004;155:304–306.
25. Kaku Y, Sarai A, Murakami Y. Genetic reclassification of porcine enterovirus. J Gen Virol. 2001;82:417–424.
*26. National Pork Board. A Producers Guide to Managing PCVAD. Available at http://www.pork.org/PorkScience/Documents/PCVADBrochure.pdf. Accessed 2 May 2007.
* Non-refereed references.