Case report |
Peer reviewed |
Restriction fragment length polymorphism analysis following unexpected Mycoplasma hyopneumoniae diagnosis in a gilt multiplication herd
Análisis de muestras mediante la técnica de polimorfismo de longitud de fragmentos de restricción después de un inesperado diagnóstico a Mycoplasma hyopneumoniae en un hato de multiplicación de primerizas
Analyse du polymorphisme de taille des fragments de restriction suite à un diagnostic inattendu pour Mycoplasma hyopneumoniae dans un troupeau multiplicateur de cochettes
J. C. S. Harding, DVM, MSc, Diplomate ABVP
Department of Large Animal Clinical Sciences, Western College of Veterinary Medicine, University of Saskatchewan, Saskatoon, Saskatchewan, Canada. Corresponding author: Dr J. C. S. Harding, Department of Large Animal Clinical Sciences, Western College of Veterinary Medicine, University of Saskatchewan, 52 Campus Drive, Saskatoon, Saskatchewan, Canada S7N 5B4; Tel: 306-966-7070; Fax: 306-966-7159; E-mail: john.harding@usask.ca.
Cite as: Harding J C S. Restriction fragment length polymorphism analysis following unexpected Mycoplasma hyopneumoniae diagnosis in a gilt multiplication herd. J Swine Health Prod. 2008;16(5):247–253.
Also available as a PDF.
SummaryLung tissue from multiple pigs from a serologically negative gilt multiplication barn unexpectedly tested positive for Mycoplasma hyopneumoniae by polymerase chain reaction (PCR) and fluorescent antibody test. While sporadic coughing, dyspnea, and unthriftiness were noted on initial examination, the clinical presentation was atypical of mycoplasmal pneumonia. A detailed diagnostic investigation, including a follow-up visit, examination at slaughter, serological testing, additional necropsies, PCR, and restriction fragment length polymorphism analysis, confirmed the herd’s M hyopneumoniae-negative status. This report highlights the importance of a thorough clinical and diagnostic examination following an unexpected positive laboratory result. Moreover, the excellent feedback and collaboration with the diagnostic laboratory in this case facilitated the continuous improvement of laboratory services for the mutual benefit of the laboratory’s clientele. | ResumenResumen - El tejido pulmonar de varios cerdos de una granja multiplicadora de hembras de reemplazo, serológicamente negativa, inesperadamente resultó positivo a Mycoplasma hyopneumoniae mediante la prueba de reacción en cadena de la polimerasa (PCR por sus siglas en inglés) y a la prueba de anticuerpos fluorescentes. A pesar de que en el examen inicial se detectó tos esporádica, disnea y desgaste, la presentación clínica fue atípica a la pulmonía producida por Mycoplasma. Una investigación diagnóstica detallada, incluyendo una visita de seguimiento a la granja, examen en matadero, análisis serológicos, necropsias adicionales, PCR, y análisis de polimorfismo de longitud de fragmentos de restricción confirmó el estado negativo a M hyopneumoniae del hato. Este reporte resalta la importancia de un examen diagnóstico y clínico profundo después de un resultado positivo de laboratorio inesperado. Además, la excelente retroalimentación y colaboración con el laboratorio de diagnóstico demostrado en este caso facilitó la mejora continua de los servicios de laboratorio para el beneficio mutuo de la clientela del laboratorio. | ResuméDe manière inattendue, du tissu pulmonaire provenant de plusieurs porcs issus d’un troupeau multiplicateur de cochettes négatif pour Mycoplasma hyopneumoniae par sérologie, s’est avéré positif pour M hyopneumoniae par réaction d’amplification en chaîne par la polymérase (PCR) et par immunofluorescence. Lors de la l’examen initial, de la toux sporadique, de la dyspnée, et un pauvre état général ont été notés, et cette présentation clinique n’était pas typique de la pneumonie à mycoplasme. Une enquête diagnostique approfondie incluant une visite de suivi, des examens au moment de l’abattage, des analyses sérologiques, des nécropsies additionnelles, des analyses par PCR, et des analyses du polymorphisme de taille des fragments de restriction ont confirmé le statu négatif pour M hyopneumoniae du troupeau. Le présent rapport met en évidence l’importance d’un examen clinique complet et des épreuves diagnostiques appropriées suite à un résultat de laboratoire positif inattendu. De plus, l’excellence de la rétroaction et de la collaboration obtenue de la part du laboratoire de diagnostic dans le présent cas a facilité l’amélioration continue des services de laboratoires pour le bénéfice mutuel de la clientèle du laboratoire. |
Keywords: swine, Mycoplasma hyopneumoniae, polymerase chain reaction, restriction fragment length polymorphism, diagnosis
Search the AASV web site
for pages with similar keywords.
Received: September 11, 2007
Accepted: March 18, 2008
Mycoplasma hyopneumoniae is a globally prevalent swine respiratory pathogen of economic significance due to its impact on growth performance, feed efficiency, mortality, and associated preventive and therapeutic costs.1 Alone or in combination with other respiratory viruses and bacteria, it causes anteroventral bronchopneumonia, most often in the grow-finish production stages, but may affect pigs of all ages, including sows and boars.1 For economic reasons, maintaining an M hyopneumoniae-free status is advantageous in swine regions where the risk of re-introduction is low.
There is evidence of genetic variation among M hyopneumoniae strains,2-7 but the biological significance of this variation in field situations is poorly understood. Two other Mycoplasma species are pathogenic to swine: Mycoplasma hyosynoviae, a cause of arthritis, and Mycoplasma hyorhinis, a cause of arthritis and fibrinous polyserositis. In contrast, Mycoplasma flocculare has been isolated from the respiratory tracts of swine1 and is generally considered nonpathogenic. In vitro experiments have demonstrated that certain strains of M flocculare adhere to and cause slight tangling of cilia, but result in significantly less ciliary damage than virulent strains of M hyopneumoniae.2
Pigs naturally infected with M hyopneumoniae alone generally develop a mild, dry, nonproductive cough within 1 to 3 weeks post infection, with maximal lesions developing and resolving by 28 and 85 days post infection, respectively.8 Co-infections with M hyopneumoniae and other swine respiratory pathogens, such as porcine reproductive and respiratory syndrome virus (PRRSV),9 swine influenza virus (SIV),10 porcine circovirus type 2 (PCV2),11 or Pasteurella multocida,12 exacerbate the severity of clinical signs and lesions. In field situations, clinical expression may ultimately depend on the number and types of respiratory co-infections, virulence of the M hyopneumoniae strain involved,5 and type of production system and flow,13 as well as on environmental influences such as pen density, indoor air quality, feed medications, and vaccination status.1
Hallmark histopathologic lesions of M hyopneumoniae infection include peribronchiolar and perivascular lymphoid hyperplasia (cuffing), debris and inflammatory cell exudate in alveoli and small airways, and edematous and enlarged bronchial and mediastinal lymph nodes, but none of these lesions are pathognomonic. Diagnosis depends on cultivation or confirmation of antigen or nucleic acid in the respiratory tract. For antigen demonstration, the fluorescent antibody test (FAT)14 or immunohistochemistry (IHC)15 assays may be used, while polymerase chain reaction (PCR)16-18 is widely available for nucleic acid demonstration. In contrast, few commercial laboratories routinely attempt M hyopneumoniae cultivation due to its slow growth and requirement for specialized media.1 All M hyopneumoniae diagnostic tests are best suited for herd diagnosis because their sensitivity and specificity are higher for herds than for individual animals. In addition, seroconversion to M hyopneumoniae occurring concurrently with characteristic clinical signs and lesions supports a herd diagnosis.
Case description
This report pertains to an unexpected M hyopneumoniae diagnosis in a 9600-head gilt multiplier finishing barn located in an isolated region in western Canada.
Farm facilities and management
The farm received 65- to 72-day-old gilt pigs (average weight 24.4 kg) twice weekly from a single nursery, which in turn received 14- to 17-day-old piglets from a single 2400-sow breeding-farrowing unit. The finishing barn consisted of eight rooms, each filled in sequential order within 2 weeks. Each room contained approximately 1200 pigs evenly distributed among 60 pens (approximately 20 animals per pen) in the order in which they came off the truck. Pens were 2.4 m � 5.4 m, providing 0.67 m2 per pig with 1.5 m of slat situated at the back of each pen. A single dry self-feeder and two water nipples were located in each pen. Pigs were phase-fed for 17 to 18 weeks until sold as F1 breeding females or as market hogs at approximately 113 kg live weight.
Farm biosecurity and health status
Biosecurity of the barn was excellent. Except for a “sister” finisher barn of the same health status that received barrow pigs from the same nursery, the nearest neighbouring pig farm was > 8 km away. In addition, strict trucking, feed delivery, and herd entry protocols were in place. The farm was closed to unscheduled visitation, and all visitors observed a 36-hour pig freedom period prior to entry. The doors were locked at all times, deterrent signage was posted, dead-stock removal bins were located 500 m from the barn, rodent baiting stations were maintained, and all transport vehicles allowed on site to move market hogs or gilts were emptied, high-pressure washed, and disinfected prior to arrival.
Diets were medicated with salinomycin (Possistac; Phibro Animal Health, Regina, Saskatchewan, Canada; 25 g per tonne) during weeks 3 and 5 post placement for control of ileitis caused by Lawsonia intracellularis. The herd was PRRSV-positive and PCV2-positive on the basis of historic serological and antigen testing, but serologically negative for M hyopneumoniae, SIV (H1N1 and H3N2), and Actinobacillus pleuropneumoniae, and inconsistently serologically positive for porcine respiratory coronavirus. The low prevalence of lung lesions and average lung scores on past examinations at slaughter supported the M hyopneumoniae-negative status of the farm. In spite of a policy of no water-soluble or parenteral treatments in the barn, mortality levels were traditionally low (eg, 3.3% for a 6-month period prior to the visit). Sixty percent of gilts in each pen were vaccinated for PRRSV (PRRS-MLV; Boehringer Ingelheim Vetmedica, Burlington, Ontario, Canada) 3 weeks post placement. In addition, all selected gilts were vaccinated pre-dispatch for Haemophilus parasuis (Suvaxyn HPS; Wyeth Animal Health, Guelph, Ontario, Canada), and erysipelas (ER Bac Plus; Pfizer Animal Health, Kirkland, Quebec, Canada).
Initial herd-health inspection
During a routine herd-health inspection on July 28, 2004, most pens were heavily manured, and a noticeable odour of ammonia was present in all rooms. Unthrifty pigs were detected in the rooms housing the most recently placed pigs. The manager indicated that during the preceding 6 months, approximately 1% of the population in each room (approximately 12 pigs per room) became “unthrifty” and were euthanized for humane reasons within 4 weeks of placement. No diagnostic testing had been undertaken because mortality levels were acceptable. Unnoticed by the farm manager and staff were a few pigs that demonstrated signs of lower respiratory tract disease, including coughing, dyspnea, and tachypnea, with or without weight loss. Four pigs with representative clinical signs, 5 to 12 weeks post placement, were euthanized by captive bolt and necropsied.
Initial diagnostic findings
Tan-coloured areas of consolidated lung in the cranial and middle lobes, but no gastrointestinal lesions, were identified grossly in all necropsied pigs. Multiple sections of lung were collected from each pig and submitted fresh and fixed in 10% buffered formalin to a regional diagnostic laboratory. Moderate to severe multifocal subacute to chronic bronchointerstitial pneumonia was diagnosed in all pigs. Lung tissue from four of four, three of four, and zero of four pigs tested IHC-positive for PCV2, PRRSV, and SIV antigens, respectively. Lung tissue from two of two and one of two pigs tested positive for M hyopneumoniae by PCR19 and FAT,20 respectively. This M hyopneumoniae diagnosis was disconcerting because the farm was a gilt multiplier with ongoing sales, it was located in a very isolated site, and it was serologically negative for M hyopneumoniae. Moreover, only sporadic coughing was evident, while widespread coughing would have been expected in a large population of naive pigs acutely infected with M hyopneumoniae. Other opportunistic bacteria, including Enterococcus species, Streptococcus species, Streptococcus suis, Escherichia coli, and Pasteurella multocida were isolated from one or more lung samples, but none were considered biologically significant.
Follow-up herd inspection
All gilt sales from the unit were suspended, and the farm was revisited 2.5 weeks after the initial visit. The objectives of the second visit were to evaluate the progression of clinical signs, to perform a broader M hyopneumoniae serologic investigation of the population, and to collect additional tissues for confirmation of M hyopneumoniae infection. No changes in the health status of the upstream herds had been observed. The placement date of each room was recorded. Most pens throughout the barn remained heavily manured. During the 5- to 10-minute observation period in each room, and after the pigs were startled with loud voices and encouraged to rise, no coughing was heard in four of the eight rooms. Sporadic coughing was detected in the remaining four rooms, ranging from a solitary coughing pig in one room, to a room with approximately 15 coughing pigs (1.25% room prevalence). The rooms in which coughing was observed housed pigs 3.5, 9.5, 11.5, and 14.5 weeks post placement. Coughing was most prevalent 9.5 weeks post placement, but one had to listen closely to detect it. A few unthrifty pigs with or without dyspnea were noted in the rooms housing pigs 3.5 and 5.5 weeks post placement. No clinical signs of respiratory disease were observed in the pigs 8 days post placement.
Follow-up diagnostic findings
Serum was harvested from 20 grower pigs from each of the two groups in which the M hyopneumoniae-positive pigs had originated on July 28. These two groups were targeted for sampling as those most likely to demonstrate seroconversion if M hyopneumoniae were present in the herd. A sample size of 20 per room was selected to detect approximately 10% prevalence with approximately 90% confidence.21 In one group (7.5 weeks post placement), no coughing was detected, and a blood sample was collected from one pig selected for convenience from every third pen. In the second group (9.5 weeks post placement), sporadic coughing was noted and 11 healthy pigs (no clinical respiratory disease), eight coughing pigs, and one unthrifty pig were sampled to establish a direct comparison between “clinical” and “non-clinical” pigs within the room. All 40 pigs sampled were ELISA-negative for M hyopneumoniae.
Four pigs, 3.5 weeks post placement, were selected for necropsy examination: one pig found dead the morning of the visit and three additional pigs demonstrating respiratory disease or poor body condition. Varying degrees (6% to 15%) of anteroventral consolidation and noncollapsed areas of lung indicative of bronchointerstitial pneumonia were identified grossly in all pigs. Fresh and formalin-fixed portions of lung, including lesional and nonlesional areas, were submitted from all pigs for routine bacterial culture, histopathology, and ancillary testing of respiratory pathogens. Diffuse interstitial pneumonia was diagnosed in three of four pigs, and multifocal mixed-inflammatory-cell (nonsuppurative) pneumonia was diagnosed in the fourth pig. Acute to subacute bronchitis and bronchiolitis were diagnosed in two of four pigs. Lung tissue from all four pigs tested FAT-negative for M hyopneumoniae and M flocculare. Lung tissue from two of two pigs tested PCR-negative for M hyopneumoniae. Negative PCR and FAT testing of lung and absence of characteristic peribronchiolar lymphoid hyperplasia suggested the absence of M hyopneumoniae infection. In spite of the airway epithelial necrosis found in two of four pigs, IHC for influenza type A virus was negative. Extensive PCV2 IHC staining was prominent in lung interstitium, peribronchiolar areas, and airway epithelium of all four pigs. Sporadic IHC staining for PRRSV was noted in the lung interstitium of all four pigs.
On August 20, 23 days after the initial visit, the lungs of 47 market hogs, examined at slaughter by an independent veterinarian, were free of gross lesions characteristic of mycoplasmal pneumonia.
Mycoplasma restriction fragment length polymorphism analysis
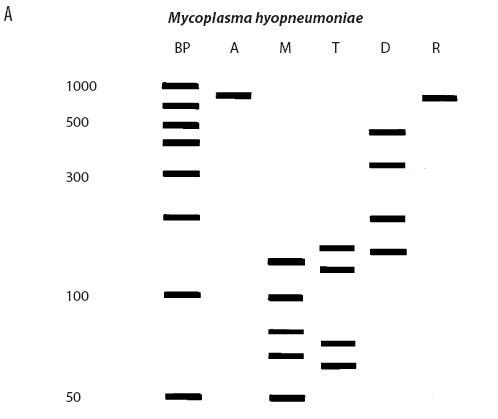
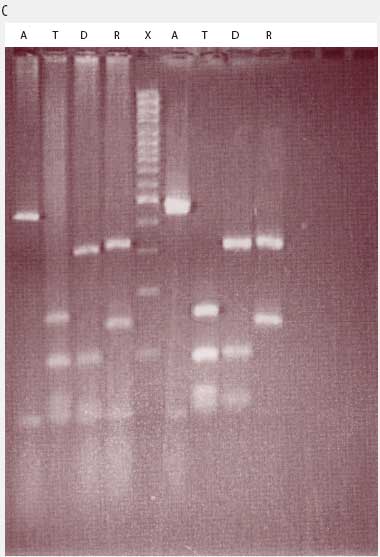
While the M hyopneumoniae PCR assay used by the laboratory19 had been satisfactorily validated and found to be very specific, the most recent slaughter-check, serological, histopathological, and PCR results strongly suggested the absence of M hyopneumoniae from the herd. To further investigate, the laboratory re-tested the frozen (-70�C) lung tissue from the two M hyopneumoniae PCR-positive pigs collected on July 28, using an alternative conventional gel-based M hyopneumoniae PCR assay (Mattsson)22 and a generic Mycoplasma species PCR followed by restriction fragment length polymorphism (RFLP) analysis.23 Briefly, the generic Mycoplasma primers amplified a region of the genome conserved by all Mycoplasma species, including those pathogenic and nonpathogenic to swine. This conserved PCR amplicon was then cut with three restriction endonucleases, DraI, RsaI (Promega, Madison, Wisconsin), and Tsp509I (New England Biolabs Inc, Ipswich, Maine), according to the instructions of the manufacturers. The resulting DNA fragment lengths were compared with the previously described DNA patterns of M flocculare, M hyorhinis, M hyosynoviae, and M hyopneumoniae.23 The results of the Mattsson PCR test were negative, and the restriction digests of both July 28 lung samples were consistent with M hyorhinis and not M hyopneumoniae (Figure 1). Samples from nine other unrelated cases diagnosed M hyopneumoniae-positive by PCR at the laboratory in 2004 were simultaneously tested using RFLP. Of these, two were M hyopneumoniae-positive, three were M hyorhinis-positive, and four had mixed mycoplasmal infections, demonstrating that the RFLP assay had the capability of detecting both M hyopneumoniae and M hyorhinis.
| Figure 1: Anticipated and actual Mycoplasma species RFLP digest patterns of (A) Mycoplasma
hyopneumoniae, (B) Mycoplasma hyorhinis, (C) Mycoplasma PCR-positive lung tissue collected from two
grower pigs (left and right of X band in panel C) demonstrating
respiratory disease and unthriftiness located in a gilt
multiplication herd. Anticipated digest patterns adapted from
Lauerman.23 Lanes: BP (panels A and B) = DNA ladder; A = Mycoplasma amplicon without restriction endonuclease
treatment; M = DNA fragments (bands) after digestion with MseI endonuclease; T = DNA fragments (bands) after digestion
with Tsp509I endonuclease; D = DNA fragments (bands) after
digestion with DraI endonuclease; R = DNA fragments (bands)
after digestion with RsaI endonuclease; X (panel C) = DNA
ladder.
|
Subsequent herd inspection and diagnostic testing
Upon receipt of the M hyopneumoniae-negative results, gilt sales were reinstated.
A subsequent visit made on October 7, 2004, indicated that the level of respiratory disease (coughing, sneezing, dyspnea) was unchanged. The number of pigs developing unthriftiness in the early grower period also remained unchanged, and total mortality followed a downward trend. At subsequent examinations at slaughter in May and November 2005, 98% and 100% percent of lungs, respectively, were free of lesions. Sera submitted for M hyopneumoniae ELISA testing (HerdChek Mycoplasma hyopneumoniae ELISA; Idexx Laboratories, Westbrook, Maine; or DAKO Mycoplasma hyopneumoniae ELISA; DAKO Corp, Carpenteria, California) on a regular basis (monthly to every other month; 220 total samples) between November 2004 and November 2007 never demonstrated a positive result.
Discussion
This report highlights the importance of a systematic diagnostic approach when investigating unexpected Mycoplasma hyopneumoniae PCR and FAT results, which could have impacted the documented health status and financial income of a gilt multiplication barn. This case also highlights the importance of collaboration between clinicians and diagnosticians in situations where unexpected results are obtained.
Various molecular methods, including amplified fragment length polymorphism, random amplified polymorphic DNA analysis, variable number of tandem repeats, sodium dodecyl sulfate polyacrylamide gel electrophoresis analysis, PCR, and RFLP have been used to compare M hyopneumoniae genotypes.5,6,24-27 In this investigation, RFLP was used to differentiate Mycoplasma species. It is not known if the wide genetic diversity of M hyopneumoniae had any impact on the RFLP results presented in this case. However, the RFLP assay detected mixed Mycoplasma infections, infection with M hyorhinis alone, and infection with M hyopneumoniae alone in lung tissue of unrelated cases that had previously tested PCR-positive for M hyopneumoniae. There is a possibility that, for technical reasons, the generalized Mycoplasma primers used for the RFLP assay failed to amplify M hyopneumoniae in the lung tissue collected on July 28, resulting in a false-negative result. This is unlikely, as the Mattsson M hyopneumoniae assay performed concurrently was also negative.
Three M hyopneumoniae ELISA assays are commercially available in North America: the Tween-20 indirect ELISA,28 the HerdChek indirect ELISA (Idexx Laboratories), and the DAKO blocking ELISA (DAKO Corp). In a recent comparison,29 all three assays had 100% specificity, but poor sensitivity (35.3% to 49.0%, depending on the ELISA).While substantially higher sensitivities (93% to 100%) are reported in other studies using experimentally8,30 and naturally infected31 pigs, it is clear that false-negative results may occur in individual pigs. However, in published reports32-38 of field investigations in nonvaccinated M hyopneumoniae-infected herds monitored using in-house ELISAs, seropositive pigs were identified in all herds. Moreover, seroconversion was first evident between 7 and 13 weeks of age, antibodies peaked between 12 and 18 weeks of age, and the seropositive prevalence was between 45% and 100%.32-38 While profiling multiple groups on a suspect herd is ideal, the prevalence of seroconversion increases with age.33,37,38 Thus, finisher pigs within 1 month of marketing are an appropriate target population for long-term serological monitoring of M hyopneumoniae-negative herds.
The case herd has remained M hyopneumoniae-seronegative for > 3 consecutive years on the basis of convenience sampling of 10 pre-dispatch gilts per month (85 kg body weight, 14 to 16 weeks post placement). A sample size of 10 animals has the power to detect 20% prevalence of seroconversion with 90% confidence.21 In spite of its potential limitations, frequent (ie, monthly) testing of smaller numbers (ie, 10 rather than 30) of late finisher animals is standard policy of some breeding companies in Canada, because it provides a practical balance between detection capability and diagnostic service costs. While isolate virulence may impact the dynamics of seroconversion,7,39 it is exceedingly unlikely that the case herd was infected with M hyopneumoniae, because the herd has remained seronegative for 3 years.
While it would have been ideal to wait a minimum of 4 weeks between the first and second visits in order to allow more time for development of a humoral immune response, it was important to minimize the farm closure period and the financial costs associated with potentially holding or slaughtering selected gilts. Moreover, if the pigs sampled on July 28 and tested by PCR and FAT had been the index pigs on the farm, the incubation period would have preceded July 28 by 2 to 3 weeks. Thus, exposure to M hyopneumoniae would have potentially occurred in early July, resulting in a minimum 5-week period from first exposure to the second visit. This would have provided sufficient time to allow for development of clinical signs and an M hyopneumoniae antibody response.
While an in-depth review of Mycoplasma PCR assays is beyond the scope of this paper, the specificity for a limited number of documented assays ranges from 93% to 100%.40-43 The sensitivity of M hyopneumoniae PCR assays is influenced by technical aspects of the assay (primer used, nested versus conventional assay), the herd’s clinical status (ie, occurrence of coughing), as well as the sample (template) tested.40-43 Since the concentration of M hyopneumoniae DNA is higher in bronchoalveolar lavage (BAL) fluid and tracheobroncheal tissue than in lung tissue and nasal swabs,41-44 the former are preferred samples for PCR. In future investigations, the submission of BAL fluid, tracheobroncheal swabs, or tissue should be considered.
The sensitivity, specificity, and positive predicted value of the M hyopneumoniae FAT are reported to be 79.0%, 97.3%, and 88.5% respectively.40 While PCR-positive, FAT-negative lung tissue may be expected, PCR-negative, FAT-positive samples are less common, but did occur in this case for reasons not fully understood.
Multiple factors, including PCV2 and PRRSV infections and poor indoor air quality, were likely associated with the interstitial pneumonia in the lungs of the pigs submitted from this barn. While PCV2 and PRRSV usually cause interstitial pneumonia, this case is unusual in that PCV2 antigen was clearly demonstrated in bronchiolar epithelium, as well as in the lung interstitium. The presence of PCV2 antigen, not type A influenza antigen, in bronchial epithelium suggests that PCV2 may have caused the bronchiolitis found in the lungs of the pigs submitted from this barn.
Although M hyorhinis is usually associated with polyserositis, lung lesions consistent with mycoplasmal pneumonia, including anteroventral consolidation and peribronchiolar and perivascular infiltration, as well as interstitial pneumonia, have been reported in experimentally inoculated pigs.45 While M hyorhinis isolates are known to be heterogeneous,46,47 the impact of strain variation on clinical presentation, virulence, or diagnostics is poorly understood. It is possible that M hyorhinis was involved in the bronchointerstitial pneumonia reported in this case, but a subclinical infection cannot be ruled out.
Implications
- Herd investigation plays a central role in accurate diagnosis of diseases on swine farms.
- Unexpected PCR-positive laboratory results should be reported to the diagnostic laboratory and further investigated by revisiting the farm and resubmitting appropriate samples for a combination of diagnostic tests if available.
- PCV2 should be included as a potential differential diagnosis in pigs with acute bronchitis and bronchiolitis.
References
1. Thacker EL. Mycoplasmal diseases. In: Straw BE, Zimmermann JJ, D’Allaire S, Taylor DJ, eds. Diseases of Swine. 9th ed. Ames, Iowa: Blackwell Publishing; 2006:701–717.
2. Young TF, Thacker EL, Erickson BZ, Ross RF. A tissue culture system to study respiratory ciliary epithelial adherence of selected swine mycoplasmas. Vet Microbiol. 2000;71:269–279.
3. Artiushin S, Minion FC. Arbitrarily primed PCR analysis of Mycoplasma hyopneumoniae field isolates demonstrates genetic heterogeneity. Int J Syst Bacteriol. 1996;46:324–328.
4. Frey J, Haldimann A, Nicolet J. Chromosomal heterogeneity of various Mycoplasma hyopneumoniae field strains. Int J Syst Bacteriol. 1992;42:275–280.
5. Vicca J, Stakenborg T, Maes D, Butaye P, Peeters J, de Kruif A, Haesebrouck F. Evaluation of virulence of Mycoplasma hyopneumoniae field isolates. Vet Microbiol. 2003;97:177–190.
6. Stakenborg T, Vicca J, Maes D, Peeters J, de Kruif A, Haesebrouck F, Butaye P. Comparison of molecular techniques for the typing of Mycoplasma hyopneumoniae isolates. J Microbiol Methods. 2006;66:263–275.
7. Meyns T, Maes D, Calus D, Ribbens S, Dewulf J, Chiers K, de Kruif A, Cox E, Decostere A, Haesebrouck F. Interactions of highly and low virulent Mycoplasma hyopneumoniae isolates with the respiratory tract of pigs. Vet Microbiol. 2007;120:87–95.
8. Sorensen V, Ahrens P, Barfod K, Feenstra AA, Feld NC, Friis NF, Bille-Hansen V, Jensen NE, Pedersen MW. Mycoplasma hyopneumoniae infection in pigs: duration of the disease and evaluation of four diagnostic assays. Vet Microbiol. 1997;54:23–34.
9. Thacker EL, Halbur PG, Ross RF, Thanawongnuwech R, Thacker BJ. Mycoplasma hyopneumoniae potentiation of porcine reproductive and respiratory syndrome virus-induced pneumonia. J Clin Microbiol. 1999;37:620–627.
10. Yazawa S, Okada M, Ono M, Fujii S, Okuda Y, Shibata I, Kida H. Experimental dual infection of pigs with an H1N1 swine influenza virus (A/Sw/Hok/2/81) and Mycoplasma hyopneumoniae. Vet Microbiol. 2004;98:221–228.
11. Opriessnig T, Thacker EL, Yu S, Fenaux M, Meng XJ, Halbur PG. Experimental reproduction of postweaning multisystemic wasting syndrome in pigs by dual infection with Mycoplasma hyopneumoniae and porcine circovirus type 2. Vet Pathol. 2004;41:624–640.
12. Amass SF, Clark LK, van Alstine WG, Bowerstock TL, Murphy DA, Knox KE, Albregts SR. Interaction of Mycoplasma hyopneumoniae and Pasteurella multocida infections in swine. JAVMA. 1994;204:102–107.
13. Sibila M, Calsamiglia M, Vidal D, Badiella L, Aldaz A, Jensen JC. Dynamics of Mycoplasma hyopneumoniae infection in 12 farms with different production systems. Can J Vet Res. 2004;68:12–18.
14. Amanfu W, Weng CN, Ross RF, Barnes HJ. Diagnosis of mycoplasmal pneumonia of swine: sequential study by direct immunofluorescence. Am J Vet Res. 1984;45:1349–1352.
15. Doster AR, Lin BC. Identification of Mycoplasma hyopneumoniae in formalin-fixed porcine lung, using an indirect immunoperoxidase method. Am J Vet Res. 1988;49:1719–1721.
16. Stakenborg T, Vicca J, Butaye P, Imberechts H, Peeters J, de Kruif A, Haesebrouck F, Maes D. A multiplex PCR to identify porcine mycoplasmas present in broth cultures. Vet Res Commun. 2006;30:239–247.
17. Calsamiglia M, Pijoan C, Trigo A. Application of a nested polymerase chain reaction assay to detect Mycoplasma hyopneumoniae from nasal swabs. J Vet Diagn Invest. 1999;11:246–251.
18. Artiushin S, Stipkovits L, Minion FC. Development of polymerase chain reaction primers to detect Mycoplasma hyopneumoniae. Mol Cell Probes. 1993;7:381–385.
19. Stemke GW, Phan R, Young TF, Ross RF. Differentiation of Mycoplasma hyopneumoniae, M flocculare, and M hyorhinis on the basis of amplification of a 16S rRNA gene sequence. Am J Vet Res. 1994;55:81–84.
20. Armstrong CH. Porcine mycoplasmas. In: Whitford HW, Rosenbusch RF, Lauerman LH, eds. Mycoplasmosis in Animals: Laboratory Diagnosis. Ames, Iowa: Iowa State University Press; 1994:70–71.
21. Pointon AM, Davies PR, Bahnson PB. Disease surveillance at slaughter. In: Straw BE, D’Allaire S, Mengeling WL, Taylor DJ, eds. Diseases of Swine. 8th ed. Ames, Iowa: Iowa State University Press; 1999:1111–1132.
22. Mattsson JG, Bergstrom K, Wallgren P, Johansson KE. Detection of Mycoplasma hyopneumoniae in nose swabs from pigs by in vitro amplification of the 16S rRNA gene. J Clin Microbiol. 1995;33:893–897.
23. Lauerman LH. Mycoplasma identification using the amplicon of a general Mycoplasma polymerase chain reaction and restriction fragment length polymorphism analysis. In: Lauerman LH, ed. Nucleic Acid Amplification Assays for Diagnosis of Animal Disease. Tukock, California: American Association of Veterinary Laboratory Diagnosticians; 1998:40–94.
24. Mayor D, Zeeh F, Frey J, Kuhnert P. Diversity of Mycoplasma hyopneumoniae in pig farms revealed by direct molecular typing of clinical material. Vet Res. 2007;38:391–398.
25. de Castro LA, Pedroso TR, Kuchiishi SS, Ramenzoni M, Kich JD, Zaha A, Vainstein MH, Ferreira HB. Variable number of tandem aminoacid repeats in adhesion-related CDS products in Mycoplasma hyopneumoniae strains. Vet Microbiol. 2006;116:258–269.
26. Assuncao P, De la Fe C, Ramirez AS, Gonzalez Llamazares O, Poveda JB. Protein and antigenic variability among Mycoplasma hyopneumoniae strains by SDS-PAGE and immunoblot. Vet Res Commun. 2005;29:563–574.
27. Scarman AL, Chin JC, Eamens GJ, Delaney SF, Djordjevic SP. Identification of novel species-specific antigens of Mycoplasma hyopneumoniae by preparative SDS-PAGE ELISA profiling. Microbiology. 1997;143:663–673.
28. Bereiter M, Young TF, Joo HS, Ross RF. Evaluation of the ELISA and comparison to the complement fixation test and radial immunodiffusion enzyme assay for detection of antibodies against Mycoplasma hyopneumoniae in swine serum. Vet Microbiol. 1990;25:177–192.
29. Erlandson KR, Evans RB, Thacker BJ, Wegner MW, Thacker EL. Evaluation of three serum antibody enzyme-linked immunosorbent assays for Mycoplasma hyopneumoniae. J Swine Health Prod. 2005;13:198–203.
30. Ameri-Mahabadi M, Zhou EM, Hsu WH. Comparison of two swine Mycoplasma hyopneumoniae enzyme-linked immunosorbent assays for detection of antibodies from vaccinated pigs and field serum samples. J Vet Diagn Invest. 2005;17:61–64.
31. Sorensen V, Barford K, Feld NC. Evaluation of a monoclonal blocking ELISA and IHA for antibodies to Mycoplasma hyopneumoniae in SPF-pig herds. Vet Rec. 1992;130:488–490.
32. Kim H, Kim E, Moon H, Kim T, Lim J, Lee Y, Park B. Analysis of Mycoplasma hyopneumoniae infection by seroepidemiological investigation in Korean swine herds. Korean J Vet Res. 2004;44:587–591.
33. Vicca J, Maes D, Thermote L, Peeters J, Haesebrouck F, de Kruif A. Patterns of Mycoplasma hyopneumoniae infections in Belgian farrow-to-finish pig herds with diverging disease-course. J Vet Med B. 2002;49:349–353.
34. Garlaite K, Siugzdaite J, Siugzda R. Vaccination against swine enzootic pneumonia under field conditions in different housing and environmental conditions. Acta Veterinaria (Beograd). 2004;54:43–52.
35. Siugzdaite J, Garlaite K, Urbsiene D. Evaluation of antibody formation, daily weight gain and meat quality after vaccination of piglets against Mycoplasma hyopneumoniae. Acta Veterinaria Hungarica. 2003;51:273–281.
36. Llopart D, Casal J, Clota J, Navarra I, March R, Riera P, Artigas C. Evaluation of the field efficacy and safety of a Mycoplasma hyopneumoniae vaccine in finishing pigs. Pig J. 2002;49:70–83.
37. Sibila M, Nofrarias M, Lopez-Soria S, Segales J, Valero O, Espinal A, Calsamiglia M. Chronological study of Mycoplasma hyopneumoniae infection, seroconversion and associated lung lesions in vaccinated and non-vaccinated pigs. Vet Microbiol. 2007;122:97–107.
38. Seok H, Joo H. Investigation of seroepidemiology of Mycoplasma hyopneumoniae infection and establishment of an on-farm eradication protocol. Korean J Vet Res. 1999;39:1218–1223.
39. Meyns T, Maes D, Dewulf J, Vicca J, Haesebrouck F, de Kruif A. Quantification of the spread of Mycoplasma hyopneumoniae in nursery pigs using transmission experiments. Prev Vet Med. 2004;66:265–275.
40. Cai HY, van Dreumel T, McEwen B, Hornby G, Bell-Rogers P, McRaild P, Josephson G, Maxie G. Application and field validation of a PCR assay for the detection of Mycoplasma hyopneumoniae from swine lung tissue samples. J Vet Diagn Invest. 2007;19:91–95.
41. Caron J, Ouardani M, Dea S. Diagnosis and differentiation of Mycoplasma hyopneumoniae and Mycoplasma hyorhinis infections in pigs by PCR amplification of the p36 and p46 genes. J Clin Microbiol. 2000;38:1390–1396.
42. Zeeh F, Kuhnert P, Miserez R, Doherr MG, Zimmermann W. Field validation of a real-time PCR test for the detection of Mycoplasma hyopneumoniae in porcine nasal swabs of live pigs. Schweizer Archiv fur Tierheilkunde. 2005;147:373–379.
43. Dubosson CR, Conzelmann C, Miserez R, Boerlin P, Frey J, Zimmermann W, Hani H, Kuhnert P. Development of two real-time PCR assays for the detection of Mycoplasma hyopneumoniae in clinical samples. Vet Microbiol. 2004;102:55–65.
44. Kurth KT, Hsu T, Snook ER, Thacker EL, Thacker BJ, Minion FC. Use of a Mycoplasma hyopneumoniae nested polymerase chain reaction test to determine the optimal sampling sites in swine. J Vet Diagn Invest. 2002;14:463–469.
45. Lin JH, Chen SP, Yeh KS, Weng CN. Mycoplasma hyorhinis in Taiwan: Diagnosis and isolation of swine pneumonia pathogen. Vet Microbiol. 2006;115:111-116.
46. Citti C, Watson-McKown R, Droesse M, Wise KS. Gene families encoding phase- and size-variable surface lipoproteins of Mycoplasma hyorhinis. J Bacteriol. 2000;182:1356–1363.
47. Rosengarten R, Wise KS. The Vlp system of Mycoplasma hyorhinis: combinatorial expression of distinct size variant lipoproteins generating high-frequency surface antigenic variation. J Bacteriol. 1991;173:4782–4793.