| Brief Communication | Peer reviewed |
Cite as: Verhegghe M, Crombé F, De Man I, et al. Preliminary study of the effect of sow washing, as performed on the farm, on livestock-associated methicillin-resistant Staphylococcus aureus skin status and strain diversity. J Swine Health Prod. 2013;21(6):313–318.
Also available as a PDF.
SummaryWashing sows (n = 12 per herd) on four Belgian pig farms positive for methicillin-resistant Staphylococcus aureus (MRSA) had no significant effect on MRSA status of the sow’s skin (P = .32) or nares (P = 1.00). In 64% of cases, the same strain was detected before and after washing. | ResumenEl lavado de hembras (n = 12 por hato) en cuatro granjas porcinas Belgas positivas a Staphylococcus aureus resistente a la metacilina (MRSA por sus siglas en inglés) no tuvo un efecto significativo en el estatus de MRSA en la piel de la hembra (P = .32) o narinas (P = 1.00). En 64% de los casos, se detectó la misma cepa antes y después del lavado. | ResuméLe lavage des truies (n = 12 par troupeau) sur quatre fermes porcines belges positives pour la présence de Staphylococcus aureus résistant à la méthicilline (MRSA) n’avait aucun effet significatif sur le statut MRSA de la peau des truies (P = .32) ou des narines (P = 1.00). Dans 64% des cas, la même souche a été détectée avant et après le lavage. |
Keywords: swine, methicillin-resistant Staphylococcus aureus, hygienic measures, sow, animal husbandry, MRSA
Search the AASV web site
for pages with similar keywords.
Received: October 19, 2012
Accepted: May 1, 2013
In 2005, a new methicillin-resistant Staphylococcus aureus (MRSA) type was isolated from swine and swine farmers.1 This livestock-associated MRSA (LA-MRSA) is now found almost worldwide in livestock, most often in swine.2 In general, colonized animals show no signs of disease, but are considered a potential source of MRSA for the human population.3 In Europe, multilocus sequence typing has shown that the majority of LA-MRSA strains belong to clonal complex 398 (CC398).2
On infected sow farms, piglets are likely to be infected with LA-MRSA after contact with the sows, the environment, other piglets, and animal-care attendants.4 The sow’s MRSA status at farrowing significantly affects the piglet’s MRSA status.5 Therefore, a reduction in the proportion of MRSA-positive sows may reduce or postpone MRSA transmission to the piglets. At present, little is known about the effect of hygienic practices on the prevalence of MRSA in sows. In the Council Directive 2008/120/EC of the European Union,6 it is stated that pregnant sows and gilts must be thoroughly cleaned when placed in farrowing crates, which results in a clean sow that can be housed in the cleaned farrowing barn. In Belgium, sow washing is a commonly used biosecurity measure on farrow-to-finish farms before sows enter the farrowing barn or upon entry (Animal Health Care Flanders, Drongen, Belgium; oral communication, 2013). During the present study, the sow-washing procedures of four farms were studied, with the aim first to determine the effect of sow washing before or upon entering the farrowing barn on the presence of MRSA on the sow’s skin, and second, to study the MRSA strains carried by the sows before and after washing.
Materials and methods
As sampled animals were not harmed, and according to the European Convention for the Protection of Vertebrate Animals used for Experimental and Other Scientific Purposes ETS 123,7 no animal utilization protocol was needed.
To select the four farms for the study, 30 pig farms were screened for MRSA between March and June, 2009.8 Those included were farrow-to-finish farms located in Flanders that had MRSA-positive swine. The present study was performed between July 2009 and December 2010. Farms were screened by nasal sampling 10 swine on each farm using a single swab pre-moistened with salt-enriched nutrient broth. Swabs were processed as described for study samples.
Before the sows were washed, the farrowing barn was cleaned with water under high pressure after manually removing the dirt. On Farms A and B, sows were washed one at a time in the gestation barn, and then walked to the farrowing barn (approximately 30 m). On Farms C and D, sows were first transported to the farrowing barn where they were washed. Sow washing consisted of three steps (Figure 1). Briefly, sows were sprayed with water and a cleaning product was applied. On Farms A and B, the same brush was used to manually apply the product, whereas on Farms C and D, the product was applied with high pressure. The cleaning products, manufacturers, and active elements are described in Table 1. As a last step, the sows were rinsed with water.
Figure 1: Schematic overview of the sow-washing procedures used on four Belgian farrow-to-finish farms positive for methicillin-resistant Staphylococcus aureus. Information about the cleaning products used is presented in Table 1. On Farm A and Farm B, the sows were manually washed with the same brush in the gestation unit and then walked one by one to the farrowing unit. On Farm C and Farm D, the sows were washed with high pressure in the farrowing unit. 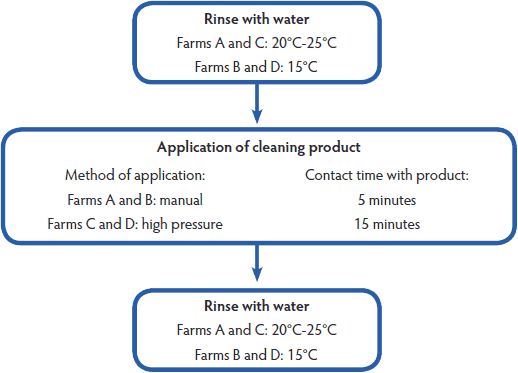 |
Table 1: Characteristics of the products used for washing sows on four Belgian pig farms*
* Farms and washing procedures described in Figure 1. |
On each farm, a sow group consisted of 36 animals. A total of 48 sows, including 12 of the 36 sows in a group on each farm, were sampled in the nasal cavity (both nares) and the skin of the back before and after washing (within 30 minutes). The nares were sampled to determine the general MRSA status of the sow, whereas the back skin was sampled to determine the effect of washing the sow. None of the owners were willing to leave a group of sows unwashed, so no control group could be sampled. A single pre-moistened swab was used to sample both nares. The swab was moistened with 3 mL of Mueller-Hinton broth (MHB; Oxoid, Basingstoke, United Kingdom), salt-enriched with 6.5% weight per volume sodium chloride (Merck, Darmstadt, Germany). One hundred cm² of a defined area of back skin (10 cm cranial to the tail in the middle of the back) was swabbed with a premoistened sponge (7 mL salt-enriched MHB added to the sponge) held in a sterilized frame (100 cm²). All samples were processed within 2 to 3 hours after sampling. The sponge samples were placed in sterile bags, and salt-enriched MHB was added to provide a 10-1 dilution. The bags containing the sponges were agitated and a tenfold dilution series was made with the salt-enriched broth to dilution 10-3. The enrichment broth dilutions and swabs were incubated at 37°C for 18 to 20 hours. One loopful of each dilution of enrichment broth was plated onto a chromogenic selective medium for MRSA (Chrom-ID MRSA; BioMerieux, Marcy l’Etoile, France). One suspect colony was purified by plating on Chrom-ID MRSA, and one suspect isolate was stored at -20°C in brain-heart infusion broth (BHI; Oxoid) supplemented with glycerol (15% weight per volume; Fisher Scientific, Leicestershire, United Kingdom) for further typing.
From each isolate, DNA was extracted according to Strandén et al9 and then stored at -20°C until further use. For MRSA confirmation, an MRSA-specific multiplex polymerase chain reaction (PCR) was used as described by Maes et al.10 A PCR specific for CC398 was performed on the obtained MRSA isolates.11 From the 91 MRSA CC398 isolates identified, 40 were arbitrarily selected. This selection contained four and seven isolates from Farm A and Farm B, respectively, and 15 and 14 isolates from Farm C and Farm D, respectively. These isolates were spa typed according to the Ridom StaphType standard procedure (http://spaserver.ridom.de). Finally, pulsed field gel electrophoresis (PFGE) was performed with the use of BstZI (Promega, Madison, Wisconsin) as a restriction enzyme.12 The obtained restriction profiles were analyzed using Bionumerics version 6.5 (Applied Maths, St-Martens-Latem, Belgium). After performing the unweighted pair group method using averages with the Dice coefficient (tolerance 1%, tolerance change 1%, and optimization 1%), a dendrogram was obtained. A cutoff value of 97% for delineation of the different pulsotypes was used.
To determine whether sow washing has an influence on the MRSA status of the nares and skin, the data were analyzed using a general estimating equation approach with the MRSA status of the sow (nares or skin) as the dependent variable, in which we accounted that the measurements from the same sow (before and after washing) were nested within a given farm and were correlated with each other. All analysis was performed in SAS 9.2 (SAS Institute Inc, Cary, North Carolina), with P values < .05 considered statistically significant.
Results
A summary of the sampling results before and after washing per farm and per sampling site is shown in Table 2. Methicillin-resistant S aureus was isolated before washing from the nares of 19 of the 48 sows (40%) and from the skin of 24 sows (50%). After washing, MRSA was found in 19 nasal samples (40%) and in 29 skin samples (60%). No differences in skin MRSA status before and after washing were observed in 31 sows. On the skin of six sows (13%), MRSA was isolated before, but not after washing. A large heterogeneity in MRSA skin status was observed between the sow populations of the four farms. On Farm A, MRSA was infrequently isolated both before and after washing. On Farm B, only a small number of samples were MRSA-positive before washing, but after washing, all nasal samples and all but two skin samples were MRSA-positive. On Farm C and Farm D, most samples were MRSA-positive before and after washing. Differences in MRSA detection on the skin and in the nares before and after washing were not significant (P = .32 and P = 1.00, respectively), with a numerically greater risk of higher MRSA isolation after washing. None of the animals displayed skin irritation after washing.
Table 2: Overview of the methicillin-resistant Staphylococcus aureus (MRSA)-status combinations obtained after nasal and skin samples were collected from 12 sows on four Belgian farrow-to-finish farms (A, B, C, and D) before and after washing sows*
* Washing protocols described in Figure 1 and Table 1. The nares were sampled with a premoistened swab, and the back skin (10 cm cranial to the tail) with a premoistened sponge. The MRSA status of a sample was determined using a chromogenic medium for MRSA and afterwards an MRSA-specific confirmation polymerase chain reaction test. Before = sampling for detection of MRSA before the sow was washed; After = sampling for detection of MRSA after the sow was washed; Neg = negative; Pos = positive. |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
All but one isolate belonged to CC398. On Farm A, Farm C, and Farm D, spa type t011 was found, whereas on Farm B spa type t034 was dominant, with only one of the selected isolates belonging to spa type t011. Pulsed field gel electrophoresis identified one pulsotype in the four isolates obtained from Farm A (Figure 2). Two pulsotypes were observed on Farm C and three on Farm B and Farm D. On each farm, one pulsotype was predominant. For 64% of isolates originating from the same location, the same pulsotype was found before and after washing. In the majority of tested sows, the same pulsotype was detected in both the skin and nasal samples.
Figure 2: Results of pulsed field gel electrophoresis testing of 40 selected methicillin-resistant Staphylococcus aureus isolates obtained from four Belgian farrow-to-finish farms enrolled in a study of the effect of washing sows (n = 48). The sow number, origin, isolation before or after washing, and spa type are shown for each isolate. Washing protocols described in Figure 1 and Table 1. PT = pulsotype; NA = not applicable. 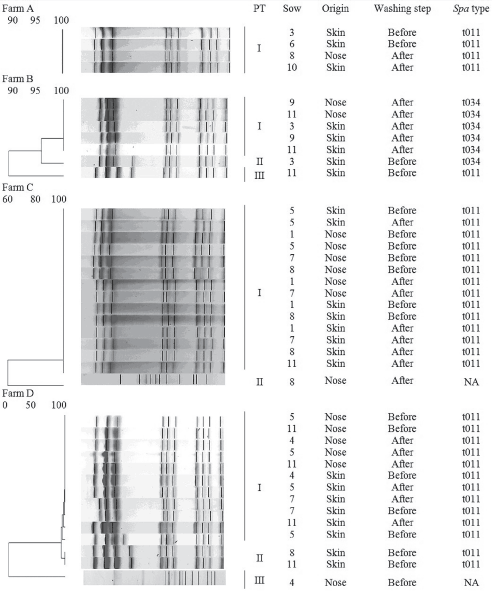 |
Discussion
Hygienic measures can help to reduce the general bacterial load of a farm, but little is known about the effect of such measures on the MRSA status of farms and animals. In Belgium, farmers commonly wash sows before they enter the farrowing barn or upon entry (Animal Health Care Flanders, Drongen, Belgium; oral communication, 2013). In a survey questionnaire, sow washing was not considered a risk factor for the presence of MRSA on a farm.13 To our knowledge, this is the first report describing the effect of sow washing, as performed on the farm, on the MRSA skin status of the sow.
Nasal samples were collected to determine the MRSA status of the sows. Methicillin-resistant S aureus is located deep within the nares and thus is thought to be uninfluenced by the washing procedure. However, the presence of MRSA in the nares might result in recolonization on the sow’s body. There was no statistically significant effect of sow washing on the sow’s skin status. We observed a numeric increase in the number of MRSA-positive samples after washing. A first explanation is that none of the washing procedures appeared sufficient to remove MRSA from colonized sows. According to the manufacturers, the active elements of Mr Clean and Fatsolve should have bacteriostatic or bactericidal activity, but little influence was observed. Since no recommended concentrations for this use were indicated, it might be interesting to determine the concentration that has an effect on a bacterial population (eg, MRSA) in vitro and subsequently test this concentration in vivo. However, caution is needed when using high concentrations of these products, which may cause skin and eye irritation, according to their Material Safety Data Sheets. Nevertheless, none of the animals sampled during the present study displayed skin irritation at the concentrations used. Control groups and more farms should be added to subsequent studies to determine the effect of the individual washing elements in the washing procedure. If subsequently the effect remains low, a disinfection step could be added to the washing procedure to reduce MRSA. In humans, a number of antiseptic products have activity against MRSA (eg, chlorhexidine, octenidine dihydrochloride, and polyhexanide), which can be evaluated in swine.14 In human medicine, one hygienic measure is often insufficient to reduce the general MRSA load in a hospital.15 So when considering decontamination of a farm, additional measures besides disinfection of the sows would most likely be required, for example, additional disinfection of the barns. However, since MRSA appears to be widespread throughout a farm, it might not be feasible to decontaminate a farm.
A second explanation for the numeric increase in MRSA-positive sows after washing may be situated in the strong bond of MRSA to corneocytes (terminally differentiated keratinocytes).16 This bond may have survived the washing process. A third explanation for the observed results might be cross-contamination during the washing process.4 On Farm B, for example, the sows were soaped manually with the same brush, possibly causing cross-contamination from an MRSA-colonized sow to a non-colonized sow. A fourth explanation might be recolonization of the animals after washing. The Farm A and Farm B sows walked one at a time to the farrowing barn after being washed in the gestation barn, which might have exposed them to an MRSA-contaminated environment (eg, the walls and floors of the corridor) or dust. In addition, when sows were being washed with warm water (Farm A and Farm C), it was noticed that the air became very humid. Spraying with water might result in the formation of aerosols in which MRSA remains present. When the animals stay in the unit, subsequent recolonization of the animals may occur. However, this hypothesis must be assessed.
All but one of the retrieved isolates belonged to CC398, which confirms the presence of LA-MRSA on these farms. Methicilllin-resistant S aureus CC398 is considered clonal, in agreement with the findings in the present study, where only one or two related spa types appeared present on a farm.5 In half of the sows, sow washing did not affect strain carriership: one MRSA strain remained throughout the procedure on each farm, which could be an indication of strain dominance as reported by Verhegghe et al.5 However, the remaining half of the sows carried different but related pulsotypes before and after washing. It is possible that sows carried multiple strains or that the dominant strains were replaced by others after washing. Since only one suspect colony per sampling event was tested by PFGE, this hypothesis still needs investigation. Therefore, the influence of sow washing on MRSA carriership could not be determined.
According to Council Directive 2008/120/EC, each farmer should clean his sows upon placing them in farrowing crates. However, in the present study, it appeared that sow washing had no effect on LA-MRSA status. On the contrary, a slight increase in MRSA isolation was observed. While this is a very small study, this result may imply that sow washing contributes to MRSA spread within a farm. The possibility exists that, in countries such as Belgium, where sow washing is often used, this measure contributes to the high prevalence of LA-MRSA. In low-prevalence countries, such as Denmark, sow washing is not a commonly used practice (Animal Health Care Flanders, Drongen, Belgium; oral communication, 2013).
In conclusion, this study describes the way sow washing was performed on four Belgian farms. This procedure did not reduce MRSA on the sow’s skin. An investigation is recommended to create an efficient and easy-to-use method to reduce the MRSA load of sows upon entry into the farrowing barn.
Implications
• Under the conditions of this preliminary study, sow washing does not reduce the presence of MRSA on the sow’s skin.
• The slight increase in MRSA isolation after washing may imply that sow washing encourages MRSA persistence within a farm.
• Since many differences in the washing procedure were observed among the four farms, further research is needed to improve and standardize the sow-washing procedure to reduce MRSA colonization.
Acknowledgements
This research was funded by the Institute for the Promotion of Innovation by Science and Technology in Flanders. We thank Rik Lenaerts for the laboratory assistance and the farmers for their collaboration on this study. Special thanks to Miriam Levenson for the English-language editing of this manuscript.
Conflict of interest
None reported.
References
1. Voss A, Loeffen F, Bakker J, Klaassen C, Wulf M. Methicillin-resistant Staphylococcus aureus in pig farming. Emerg Infect Dis. 2005;11:1965–1966.
2. Weese JS. Methicillin-resistant Staphylococcus aureus in animals. ILAR J. 2010;51:233–244.
3. Graveland H, Wagenaar JA, Bergs K, Heesterbeek H, Heederik D. Persistence of livestock associated MRSA CC398 in humans is dependent on intensity of animal contact. Plos One. 2011;6:e16830.
4. Smith TC, Moritz ED, Larson KRL, Ferguson DD. The environment as a factor in methicillin-resistant Staphylococcus aureus transmission. Rev Environ Health. 2010;25:121–134.
5. Verhegghe M, Pletinckx LJ, Crombé F, Van Weyenberg S, Haesebrouck F, Butaye P, Heyndrickx M, Rasschaert G. Cohort study for the presence of livestock-associated MRSA in piglets: Effect of sow status at farrowing and determination of the piglet colonization age. Vet Microbiol. 2013;162:679–686.
6. The Council of the European Union. Council Directive 2008/120/EC of the European Union. 2008. Available at: http://eur-lex.europa.eu/LexUriServ/LexUriServ.do?uri=OJ:L:2009:047:0005:0013:EN:PDF. Accessed 30 June 2013.
7. The European Union. 1986. European Convention for the Protection of Vertebrate Animals used for Experimental and Other Scientific Purposes. ETS No. 123. Available at: http://conventions.coe.int/treaty/en/treaties/html/123.htm. Accessed 30 June 1013.
8. Verhegghe M, Pletinckx LJ, Crombé F, Vandersmissen T, Haesebrouck F, Butaye P, Heyndrickx M, Rasschaert G. Methicillin-resistant Staphylococcus aureus (MRSA) ST398 in pig farms and multispecies farms. Zoonoses Public Health. 2013;60:366–374.
9. Strandén A, Frei R, Widmer AF. Molecular typing of methicillin-resistant Staphylococcus aureus: Can PCR replace pulsed-field gel electrophoresis? J Clin Microb. 2003;41:3181–3186.
10. Maes N, Magdalena J, Rottiers S, De Gheldre Y, Struelens MJ. Evaluation of a triplex PCR assay to discriminate Staphylococcus aureus from coagulase-negative staphylococci and determine methicillin resistance from blood cultures. J Clin Microbiol. 2002;40:1514–1517.
11. Stegger M, Liindasay JA, Moodley A, Skov R, Broens EM, Guardabassi L. Rapid PCR detection of Staphylococcus aureus clonal complex 398 by targeting the restriction-modification system carrying sau1hsdS1. J Clin Microbiol. 2011;49:732-734.
12. Rasschaert G, Vanderhaeghen W, Dewaele I, Janez N, Huijsdens X, Butaye P, Heyndrickx M. Comparison of fingerprinting methods for typing methicillin-resistant Staphylococcus aureus sequence type 398. J Clin Microbiol. 2009;47:3313–3322.
13. Broens EM, Graat EAM, Van der Wolf PJ, Van de Giessen AWV, de Jong MCM. Prevalence and risk factor analysis of livestock associated MRSA-positive pig herds in The Netherlands. Prev Vet Med. 2011;102:41–49.
14. Koburger T, Hübner NO, Braun M, Siebert J, Kramer A. Standardized comparison of antisepetic efficacy of triclosan, PVP-iodine, octinedine dihydrochloride, polyhexanide and chlorhexidine digluconate. J Antimicrob Chemother. 2010;65:1712–1719.
15. Jeanes A, Rao G, Osman M, Merrick P. Eradication of persistent environmental MRSA. J Hosp Infect. 2005;61:85–86.
16. Woolley KL, Kelly RF, Fazakerley J, Williams NJ, Nuttal TJ, McEwan NA. Reduced in vitro adherence of Staphylococcus species to feline corneocytes compared to canine and human corneocytes. Vet Dermatol. 2008;19:1–6.
